Classification rationale
1
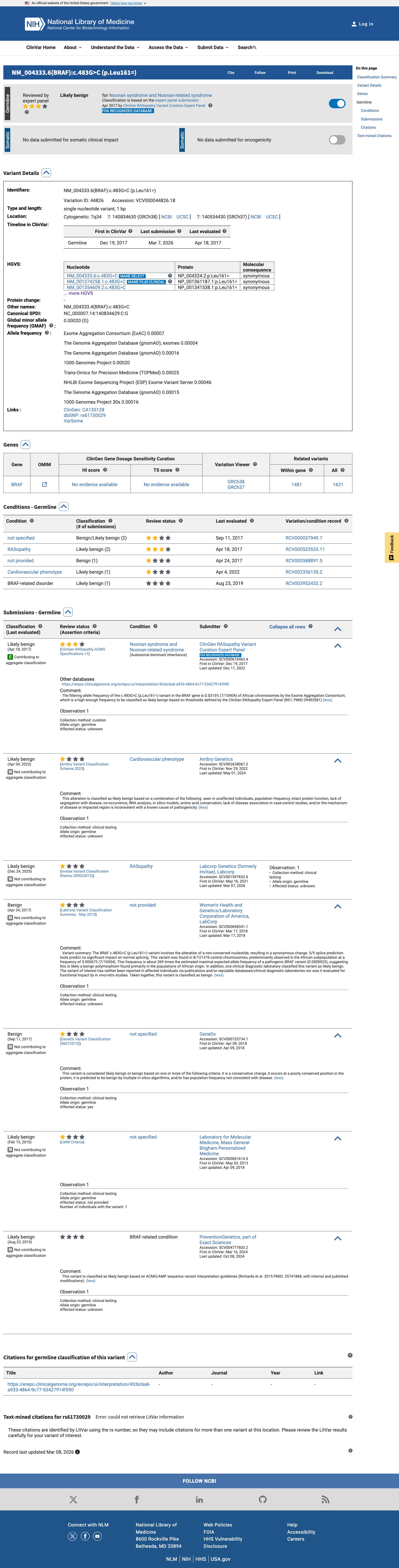
The BRAF c.483G>C (p.Leu161=) variant has not been identified as a statistically significant hotspot in Cancer Hotspots and has been reported in ClinVar as benign/likely benign, including a likely benign expert-panel classification.
hotspots ↗ clinvar ↗2
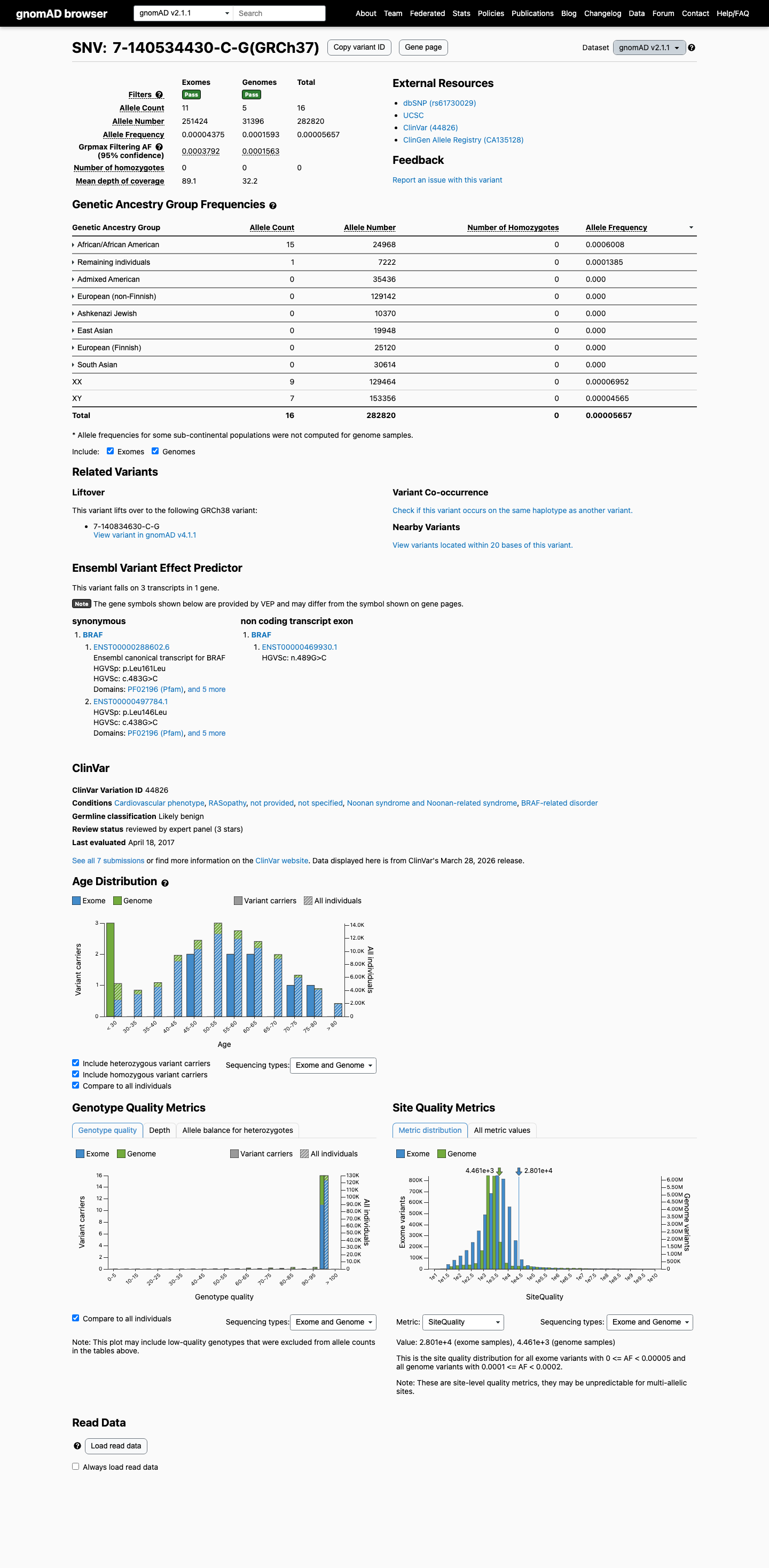
This variant is present in gnomAD, with grpmax filtering allele frequency 0.03792% in v2.1 and 0.04133% in v4.1, which is above the BRAF VCEP BS1 threshold of 0.025% but below the BA1 threshold of 0.05%.
gnomad_v2 ↗ gnomad_v4 ↗ cspec ↗3
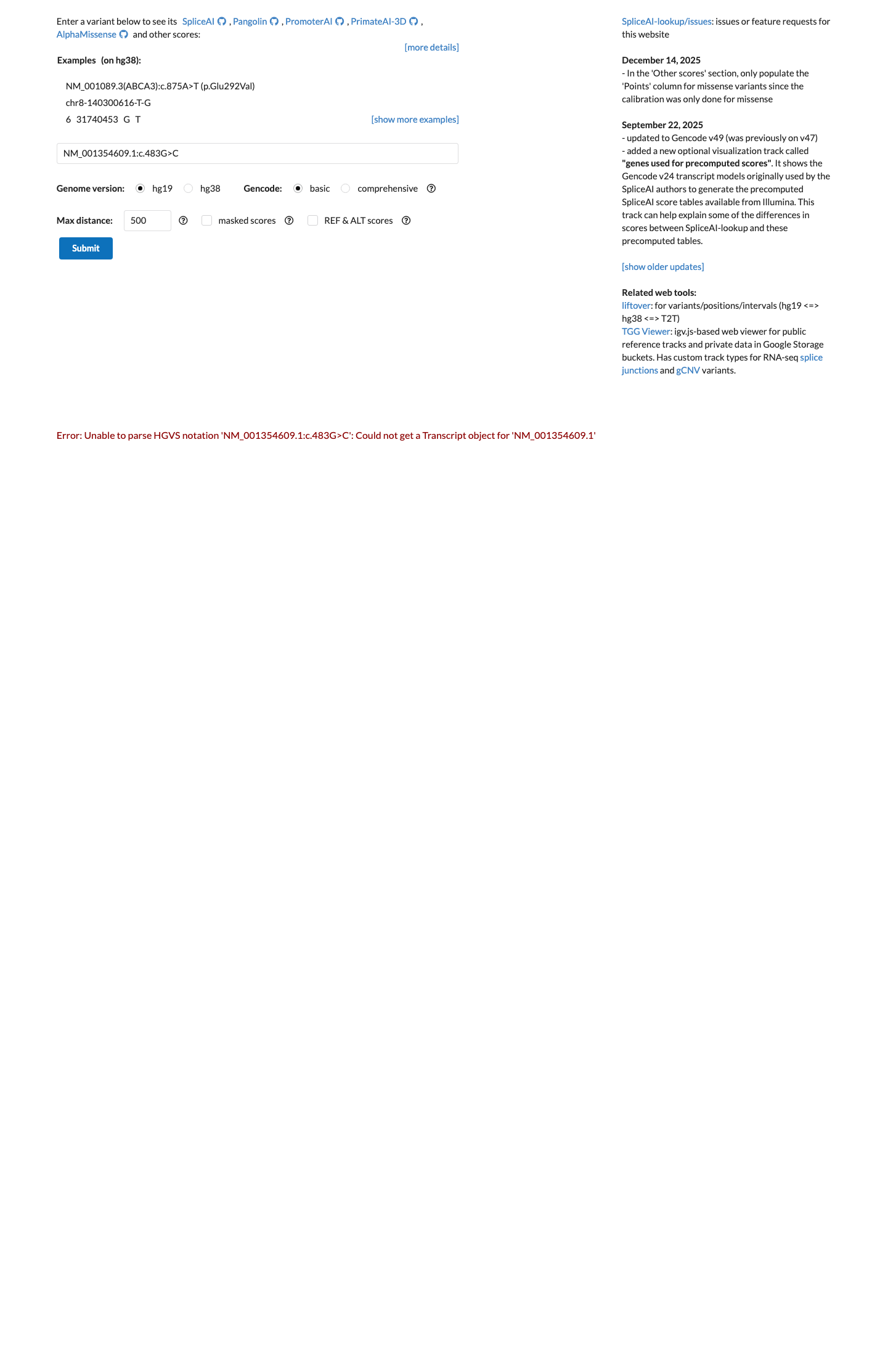
In silico splice prediction does not support a deleterious effect, with SpliceAI showing a maximum delta score of 0.02 for this synonymous change.
spliceai ↗