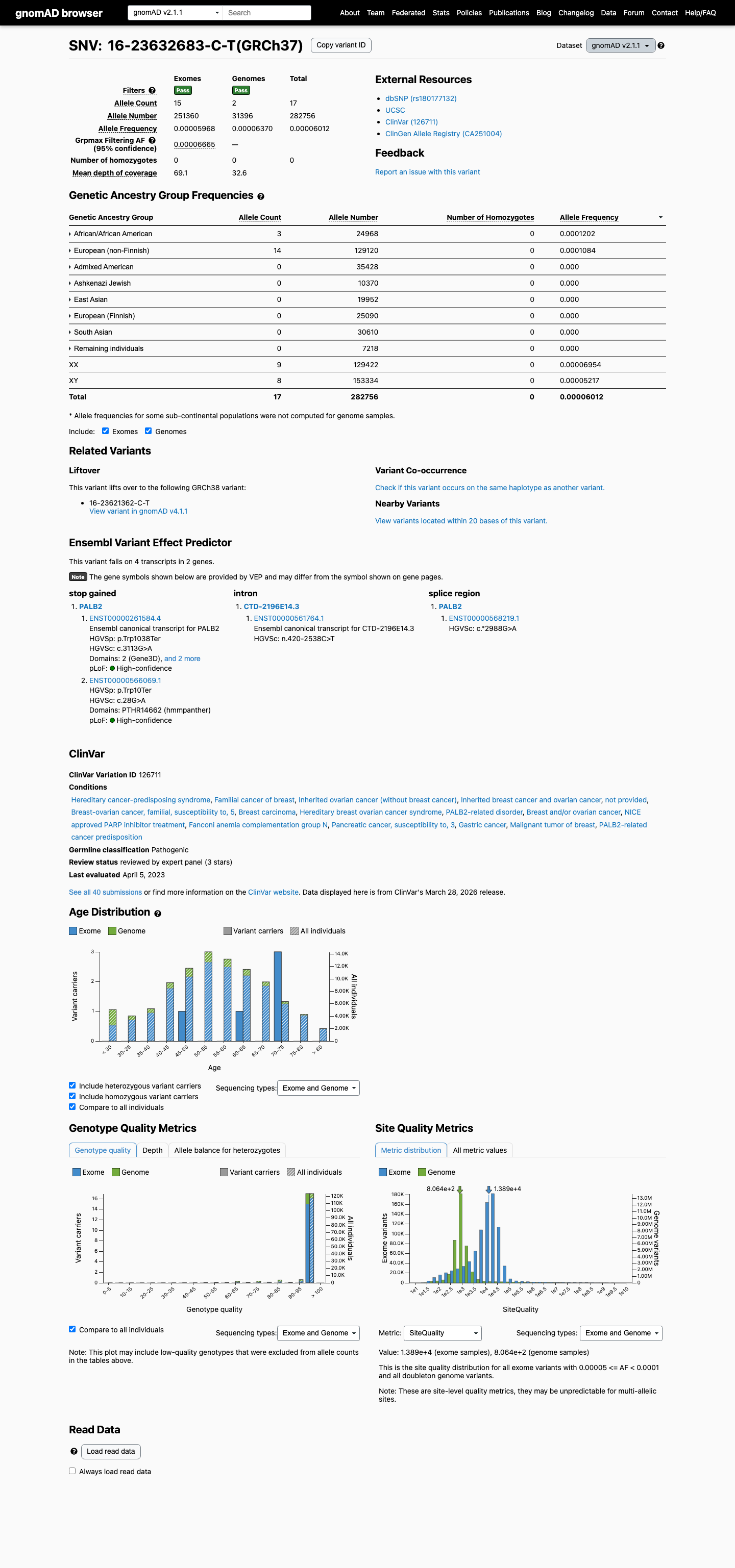
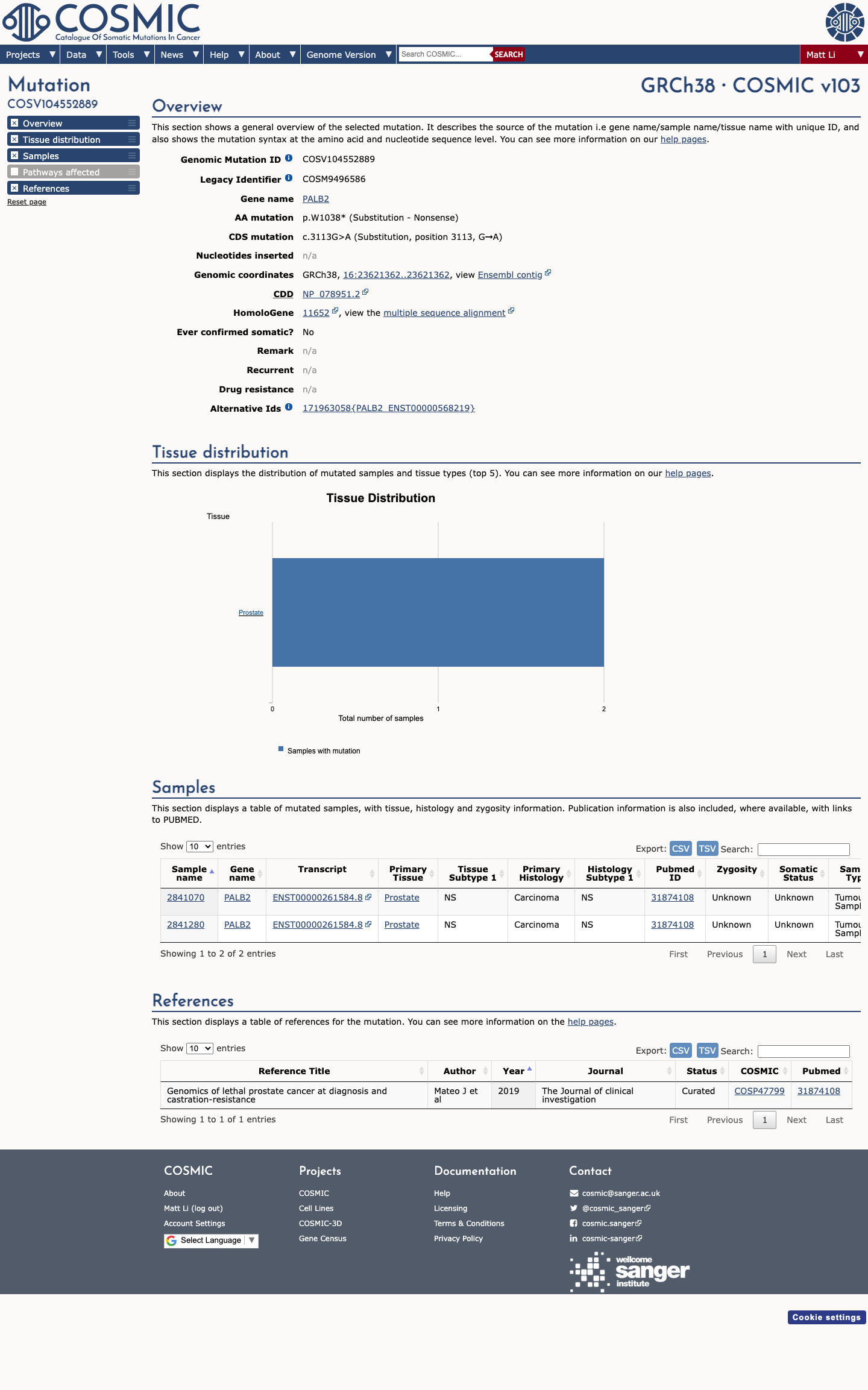
The PALB2 c.3113G>A (p.Trp1038Ter) variant has been reported in ClinVar with a pathogenic expert panel classification and additional pathogenic clinical laboratory submissions.
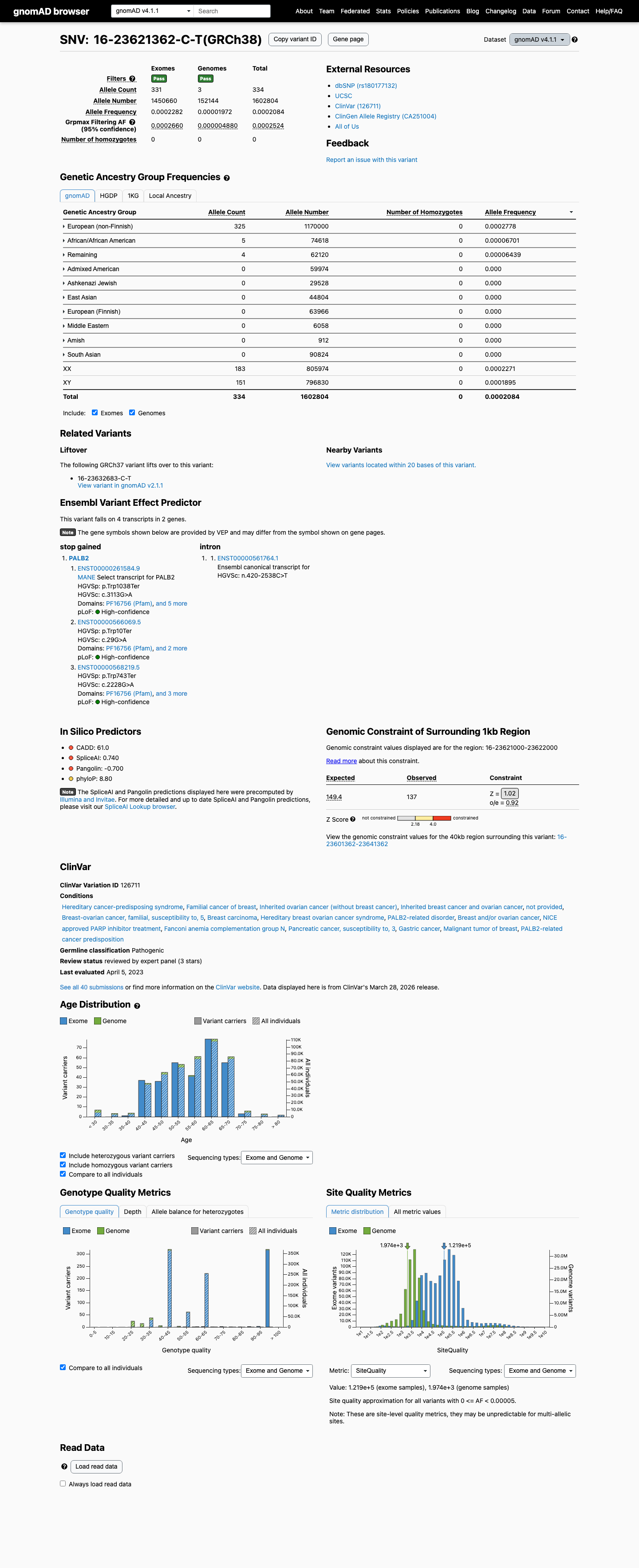
clinvar ↗This variant is present in gnomAD v4 at a total allele frequency of 0.020838% and reaches 0.027778% in the highest observed population, which is above the PALB2 BS1 threshold of 0.01% and above the PM2_Supporting threshold of 0.000333%.
gnomad_v4 ↗ cspec ↗This variant introduces a premature termination codon and is consistent with a loss-of-function effect in PALB2, a gene for which loss of function is an established disease mechanism in the PALB2 VCEP framework.
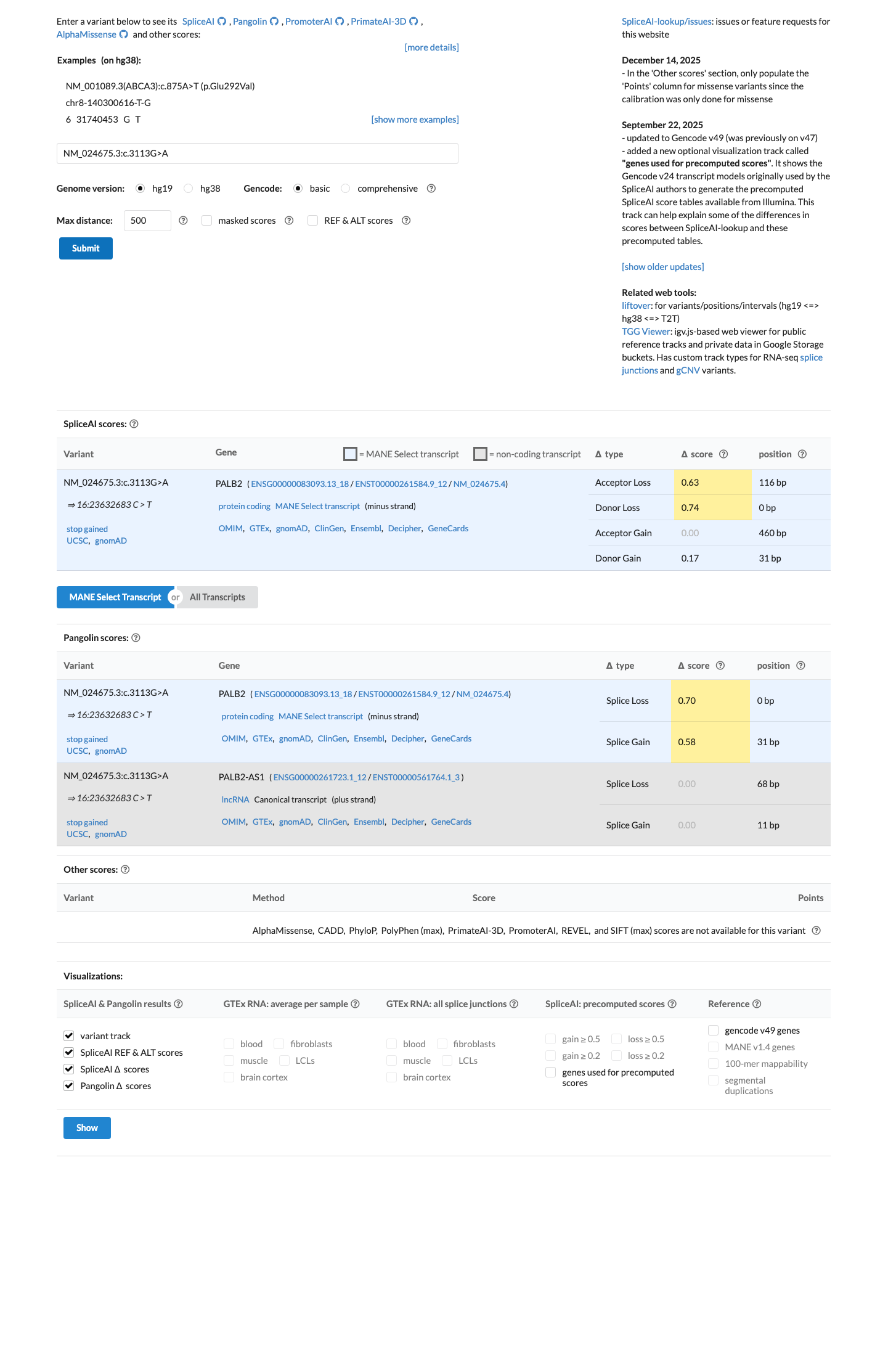
cspec ↗SpliceAI predicts splice impact with a maximum delta score of 0.74, but under PALB2-specific rules PP3 is not additionally applied for this nonsense variant type.
spliceai ↗ cspec ↗