Classification rationale
1
The CUX1 c.1657_1716+4dup (NP_001189473.1:p.?) variant has not been reported in ClinVar.
clinvar ↗2
This variant is absent from gnomAD v2.1 and gnomAD v4.1, which supports rarity in population databases.
gnomad_v2 ↗ gnomad_v4 ↗3
Germline loss of function is an established disease mechanism for CUX1, but this duplication does not fall into the generic PVS1 default null-variant categories based on the available variant-level assessment.
4
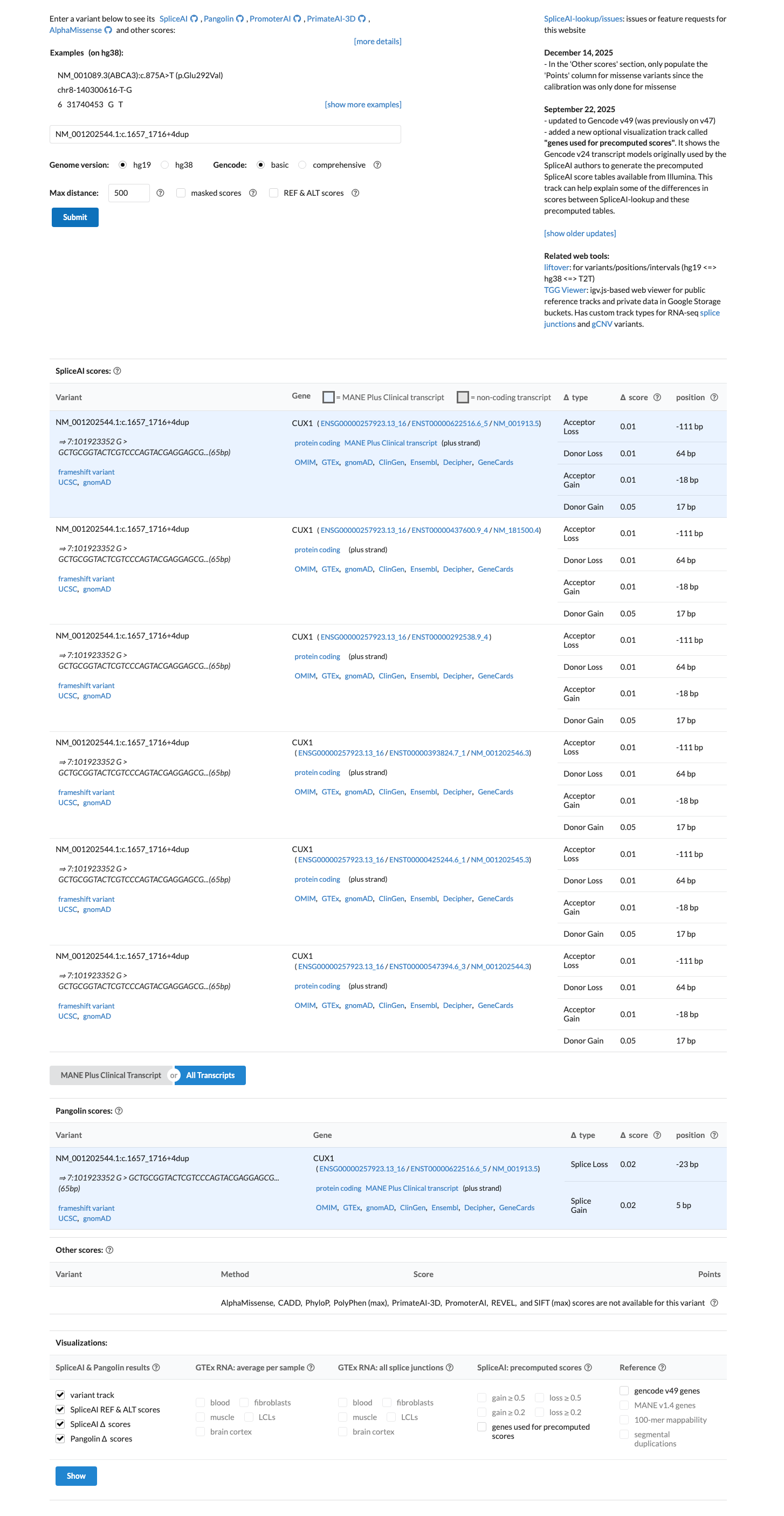
SpliceAI predicts no significant splice impact for this variant, with a maximum delta score of 0.05.
spliceai ↗