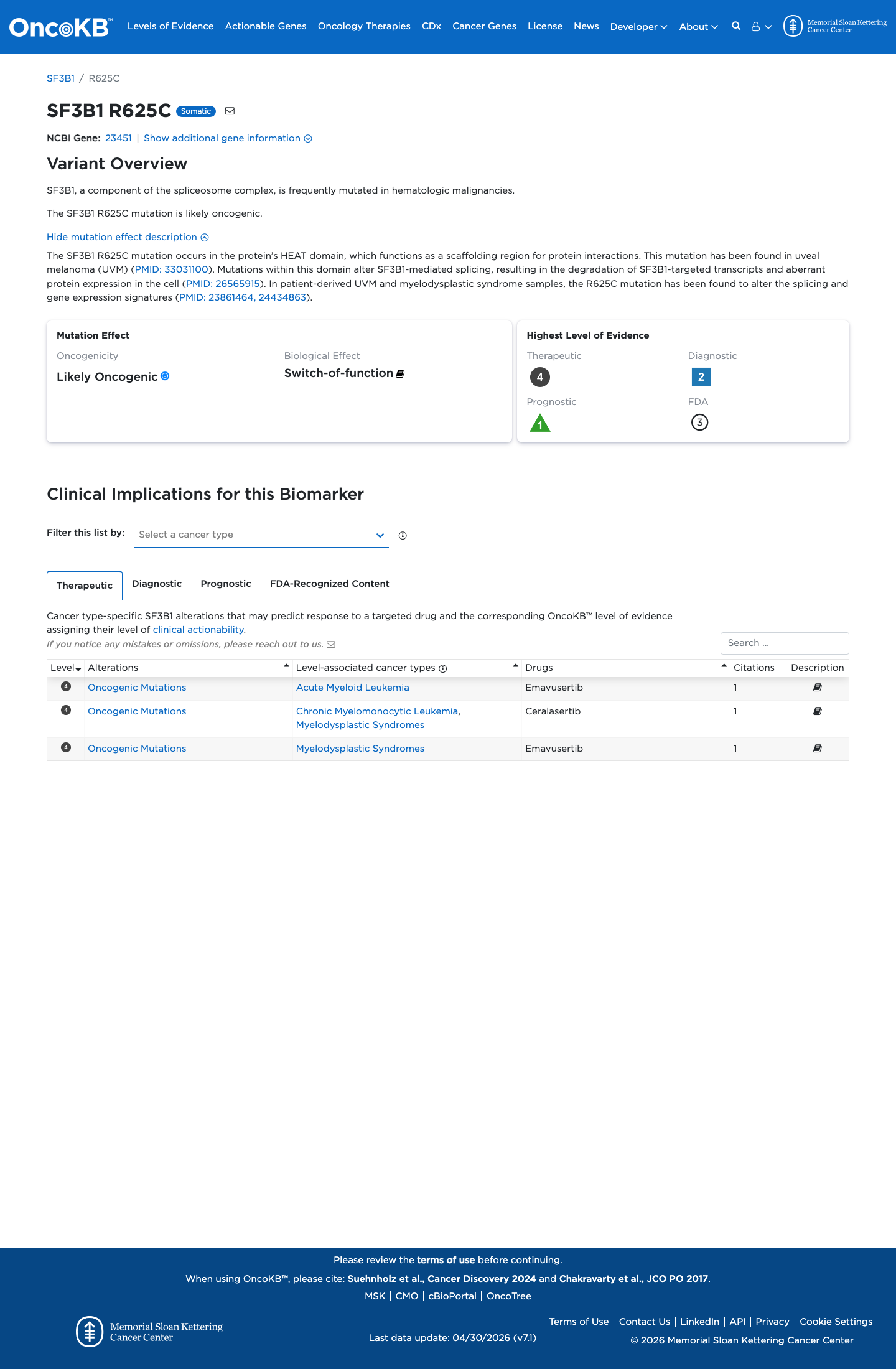
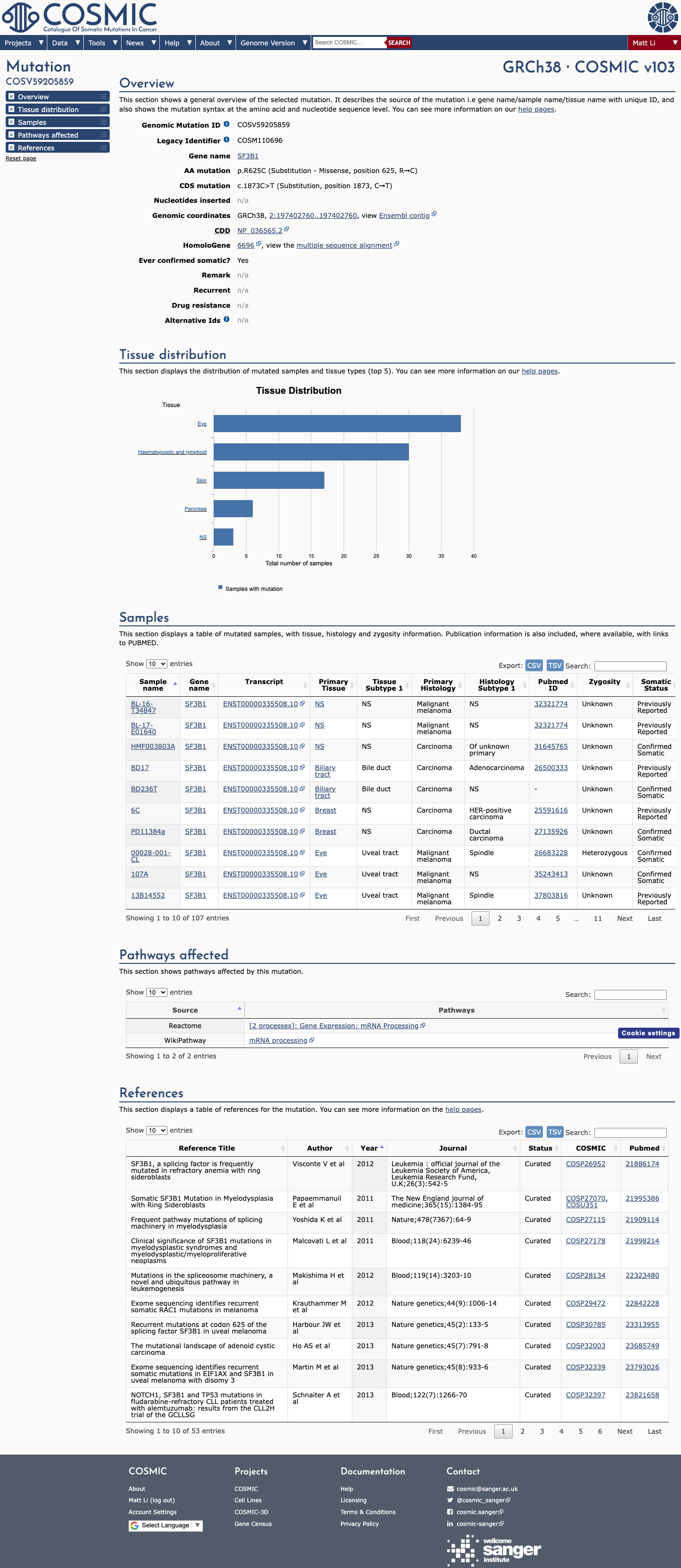
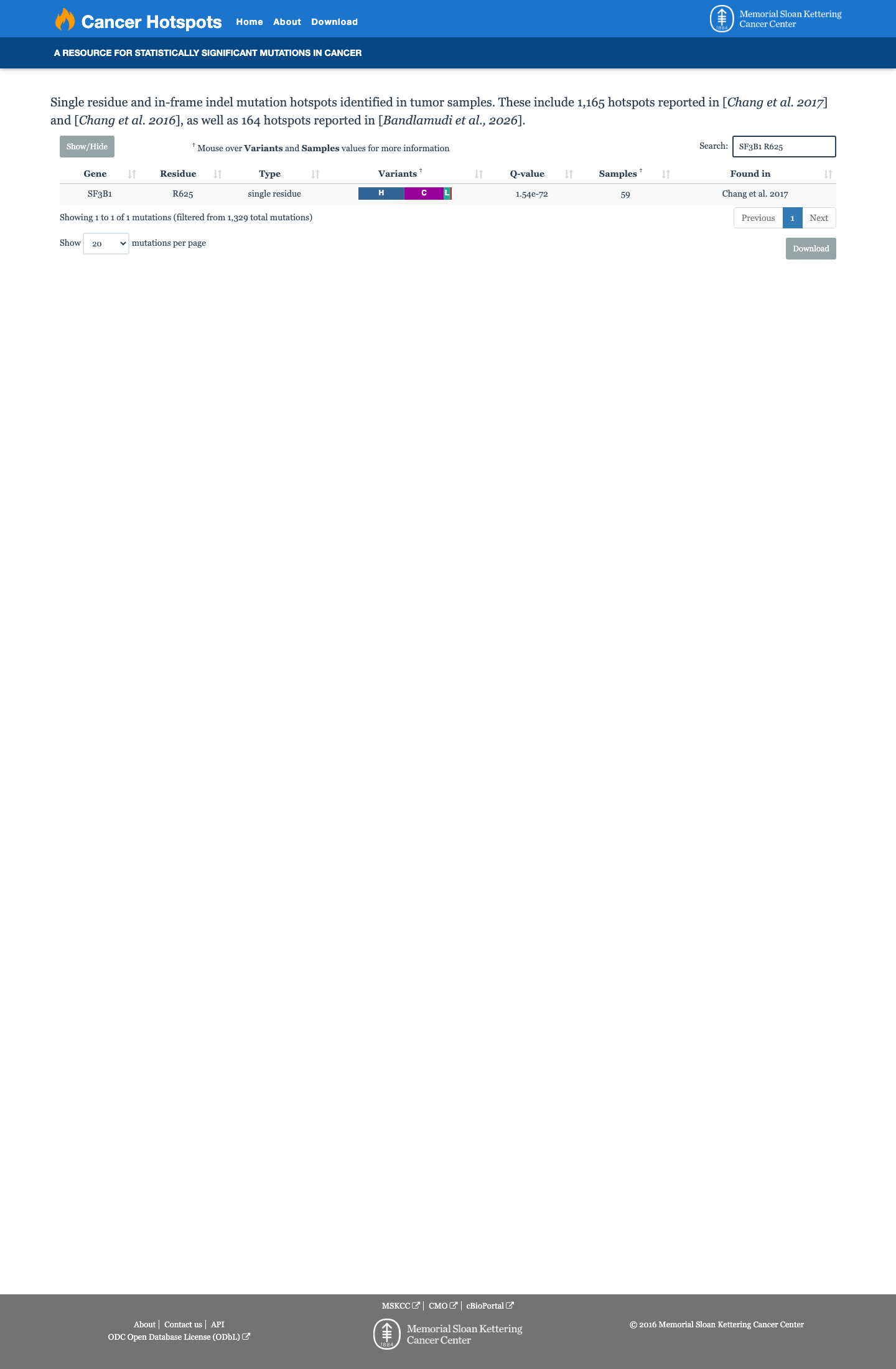
The SF3B1 c.1873C>T (p.Arg625Cys; p.R625C) variant has been observed repeatedly in somatic cancers and is recorded in ClinVar, while published studies identify codon 625 as a recurrently mutated SF3B1 hotspot.
clinvar ↗ PMID:23313955 ↗ PMID:23861464 ↗ PMID:24434863 ↗ PMID:26565915 ↗This variant is rare in population databases, with 1/251128 alleles in gnomAD v2.1 (AF 3.98203e-06; 0.00040%) and 3/1613820 alleles in gnomAD v4.1 (AF 1.85894e-06; 0.00019%), both below the 0.1% PM2 threshold.
gnomad_v2 ↗ gnomad_v4 ↗In published functional studies, SF3B1 hotspot-mutant tumors and model systems showed altered alternative splicing and abnormal 3' splice-site selection, supporting functional importance of the codon 625 region, although exact p.Arg625Cys assay-level evidence remains limited.
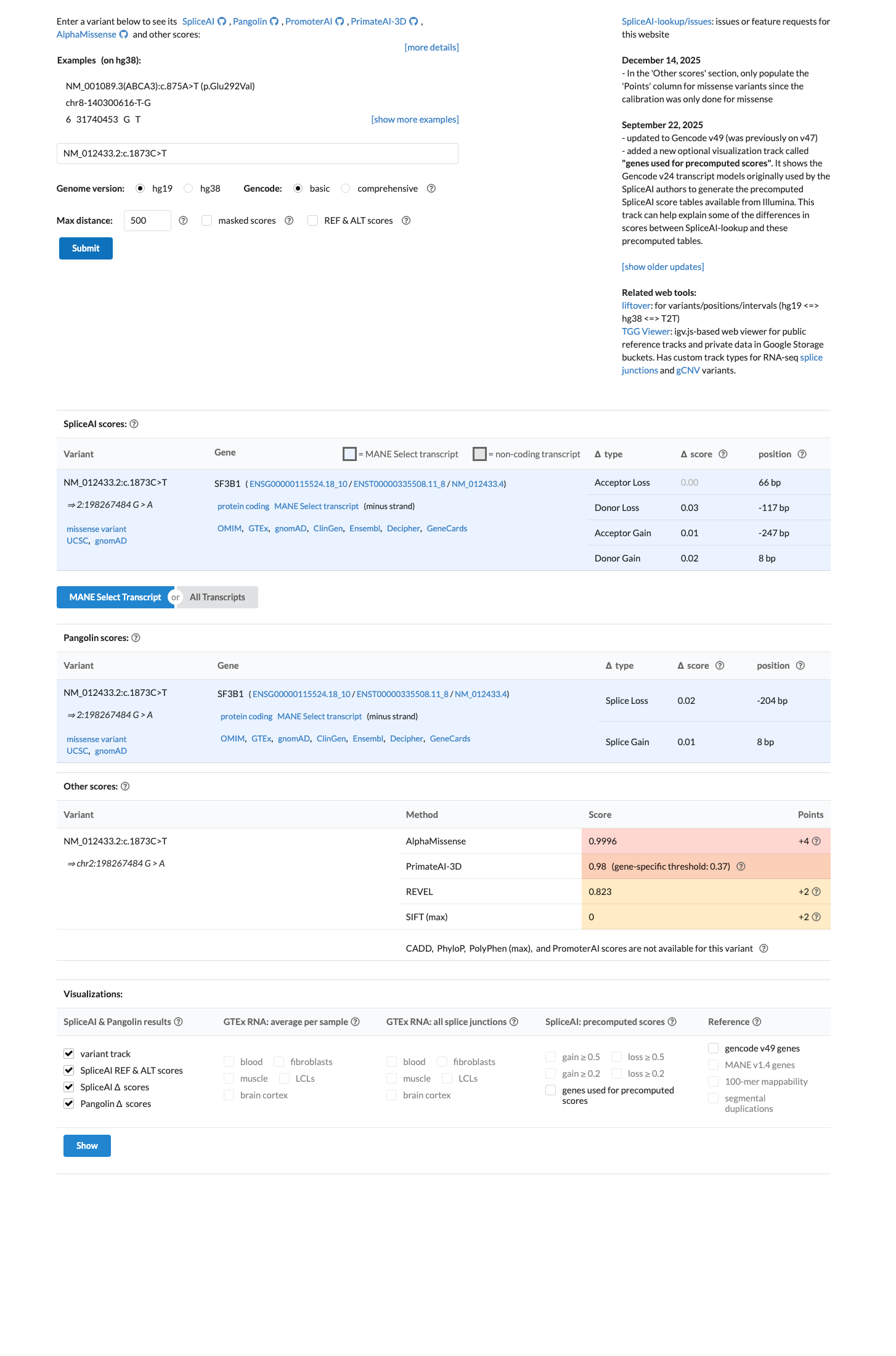
PMID:23861464 ↗ PMID:24434863 ↗ PMID:26565915 ↗ PMID:33031100 ↗Computational evidence supports a deleterious missense effect, with REVEL 0.823 and BayesDel 0.321224, while SpliceAI predicts no significant splice disruption with a maximum delta score of 0.03.
spliceai ↗