Classification rationale
1
The CHEK2 NM_007194.4:c.-6-8T>G (NP_009125.1:p.?) variant has been reported in ClinVar as likely benign by a single clinical laboratory.
clinvar ↗2
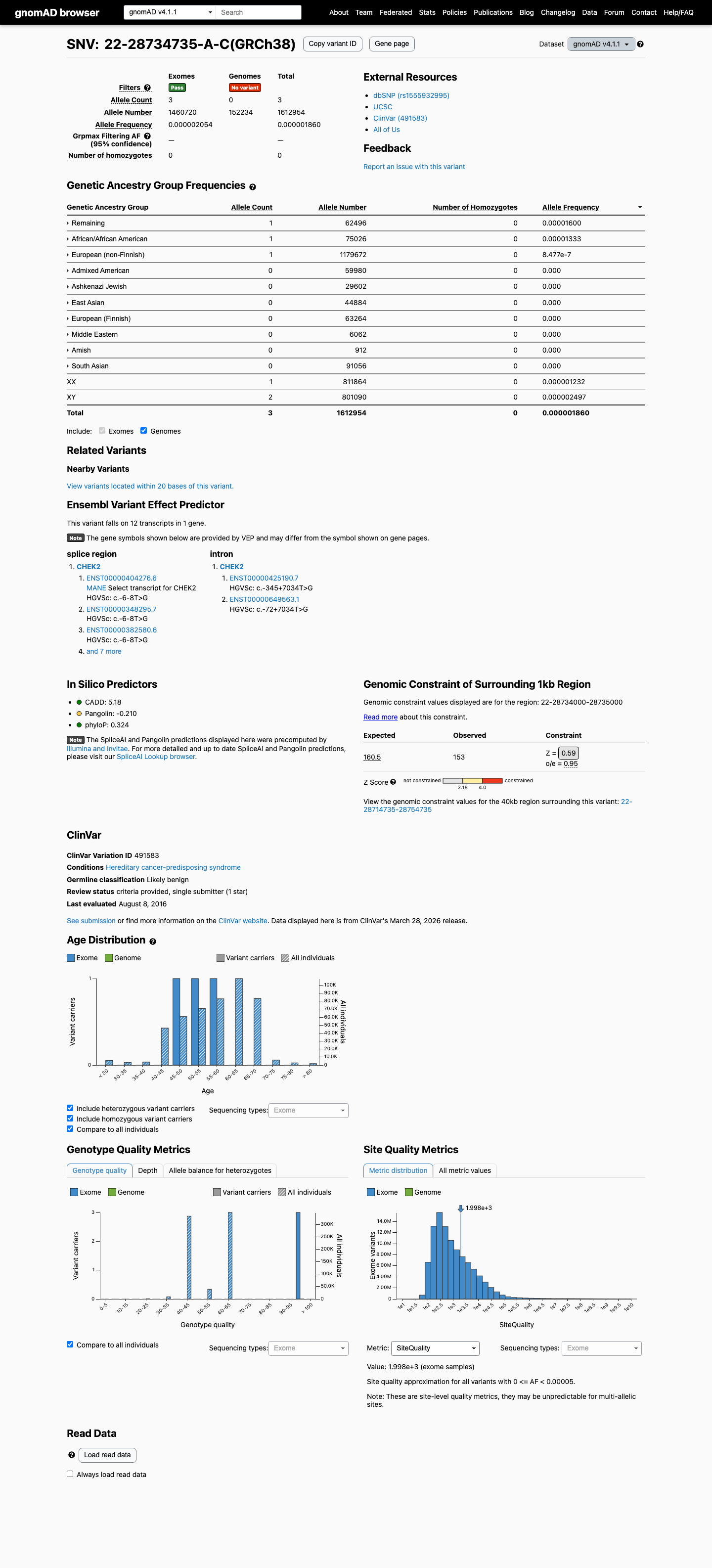
This variant is absent from gnomAD v2.1 and is present only 3 times in 1,612,954 alleles in gnomAD v4.1 (overall AF 1.85994e-06; highest observed population AF 1.6001e-05), supporting rarity but not a benign frequency threshold.
gnomad_v2 ↗ gnomad_v4 ↗3
In silico splicing analysis does not support a meaningful splice effect, with SpliceAI showing a maximum delta score of 0.10.
spliceai ↗