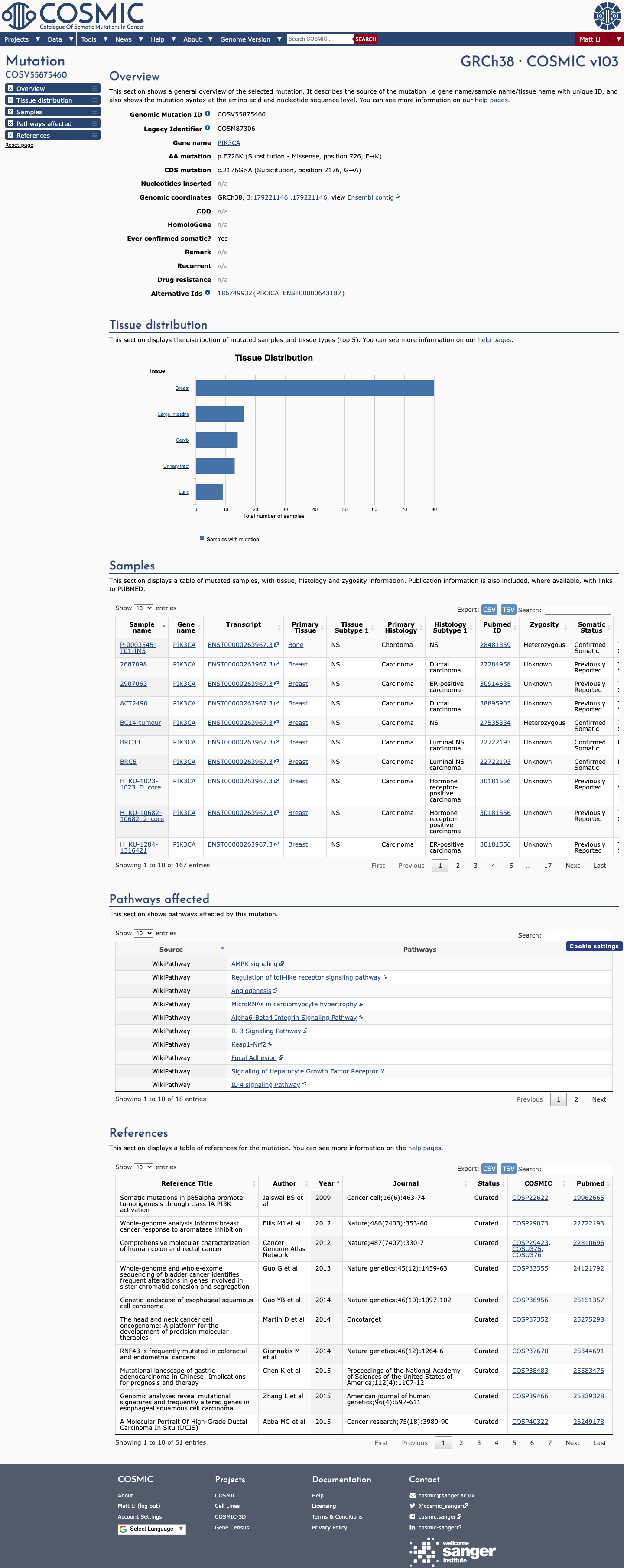
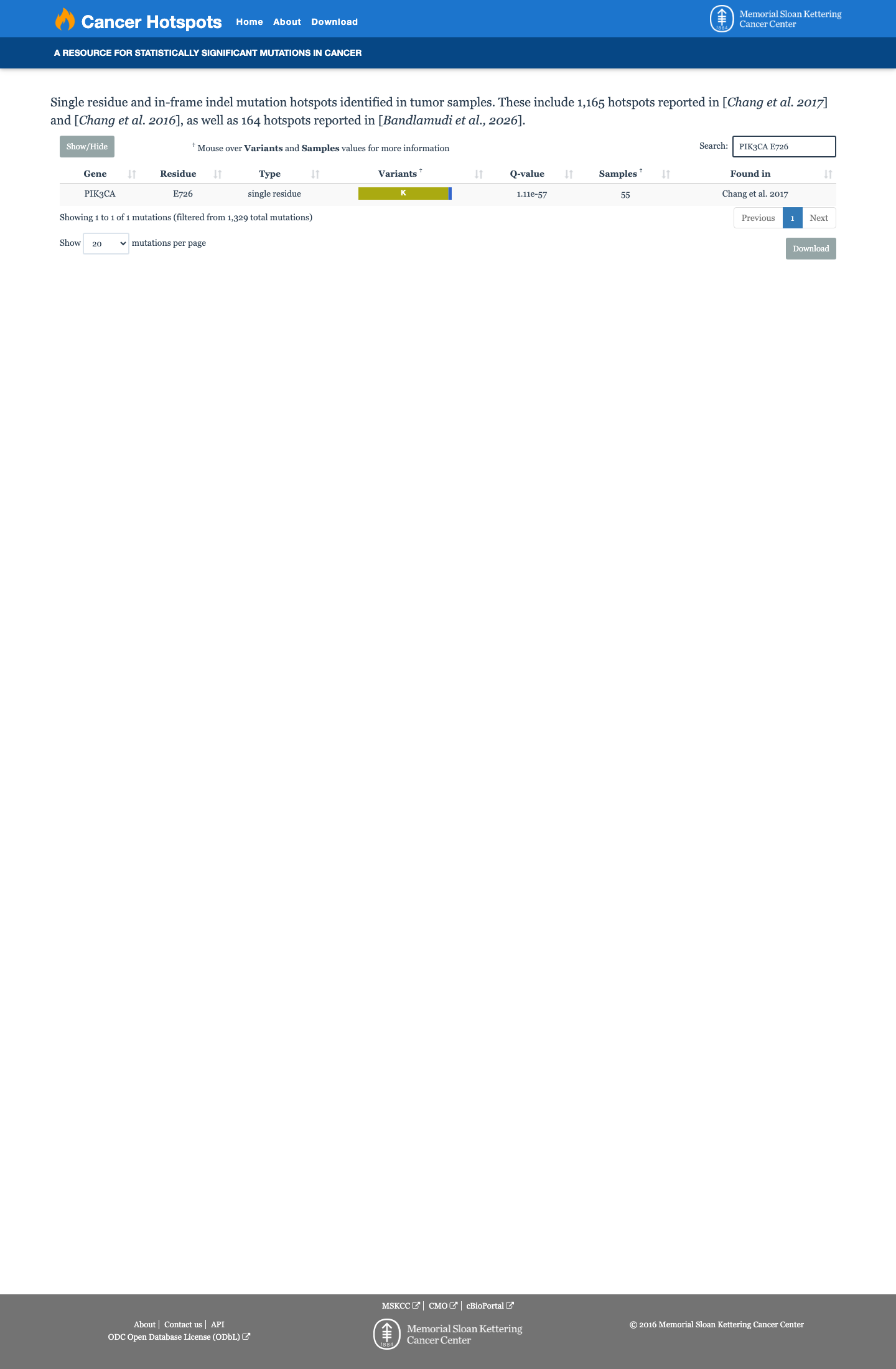
The PIK3CA c.2176G>A (p.Glu726Lys, p.E726K) variant has been observed in somatic cancers in COSMIC (COSV55875460, n=150) and has been reported in ClinVar as Pathogenic, including expert-panel review by the ClinGen Brain Malformations Variant Curation Expert Panel.
clinvar ↗This variant is absent from gnomAD v2.1 and gnomAD v4.1, which supports PM2 at Supporting strength under the Brain Malformations VCEP population rule.
gnomad_v2 ↗ gnomad_v4 ↗ cspec ↗Published PIK3CA disease-mechanism and functional literature was identified, but the retrieved materials do not provide enough variant-specific assay detail to assign PS3 or BS3 for p.Glu726Lys without direct full-text review.
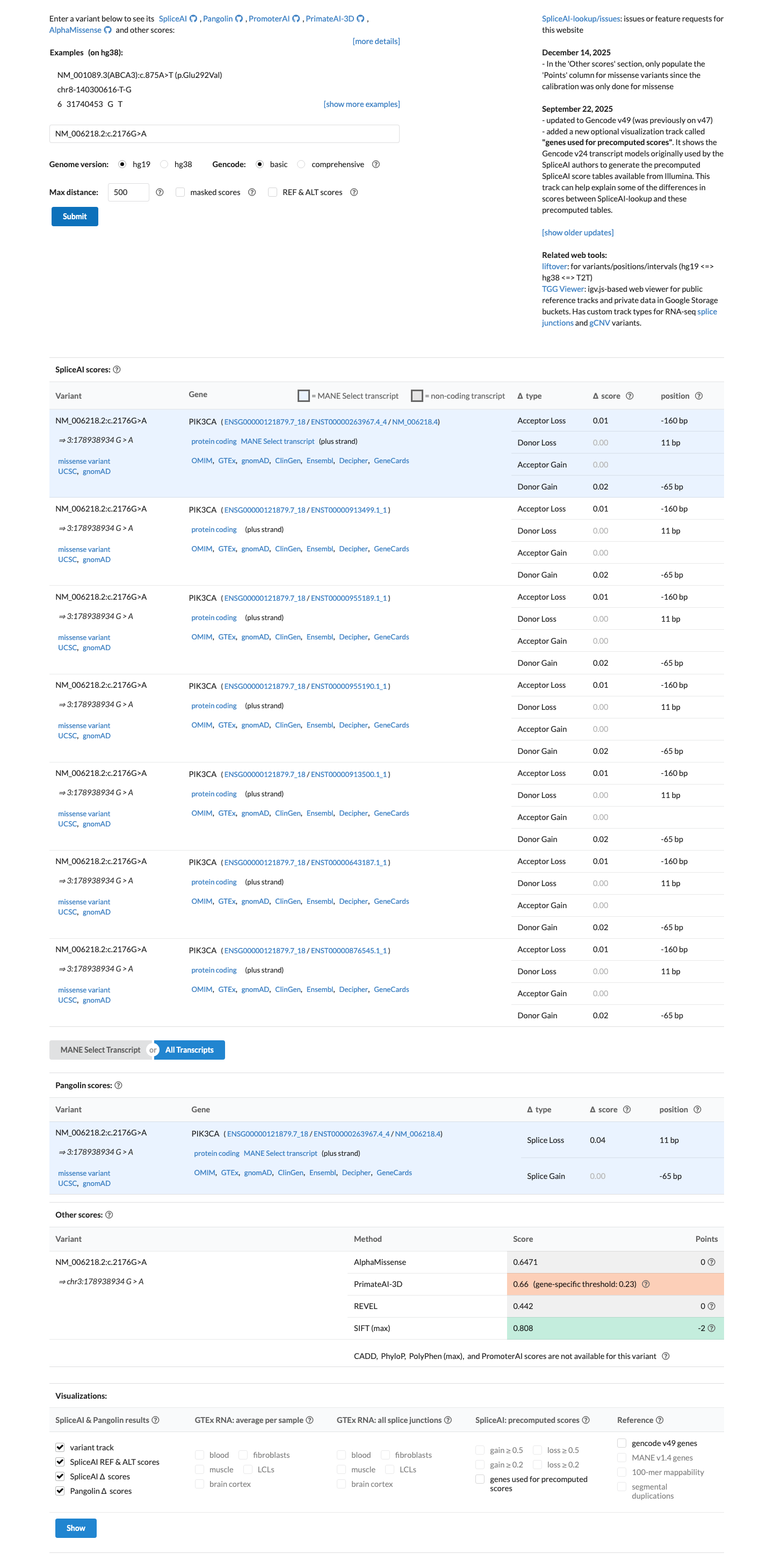
PMID:22729224 ↗ PMID:29533785 ↗ oncokb ↗In silico data show no predicted splice disruption by SpliceAI (max delta score 0.02), with REVEL 0.442 and BayesDel 0.266518; however, PP3 is not applicable for PIK3CA gain-of-function variants in this VCEP framework and BP4 is restricted to non-missense splicing-context variants.
spliceai ↗ cspec ↗