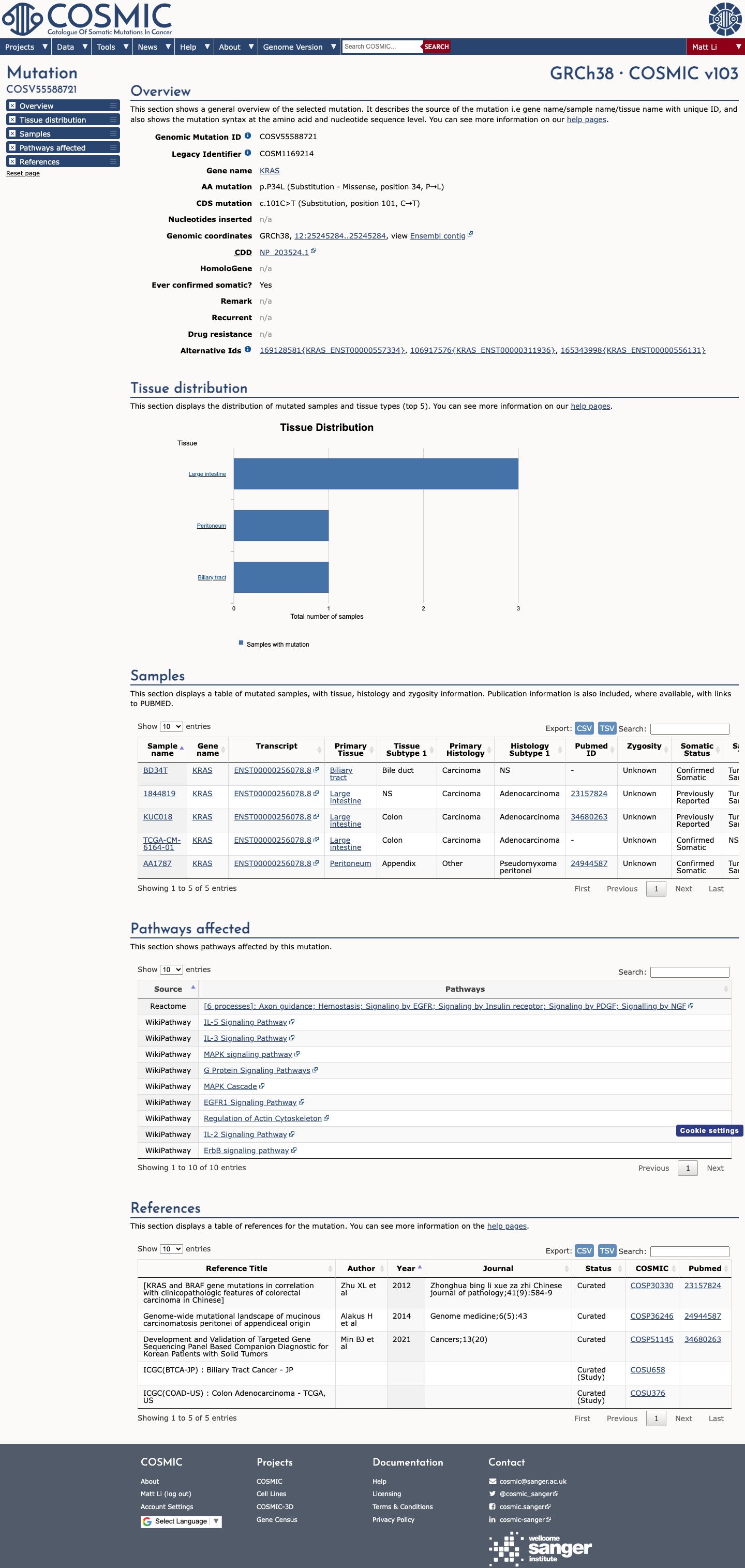
The KRAS c.101C>T (p.Pro34Leu, p.P34L) variant has been reported in ClinVar as pathogenic and has been reviewed by the ClinGen RASopathy Variant Curation Expert Panel.
clinvar ↗This variant is absent from gnomAD v2.1 and gnomAD v4.1, supporting rarity in population reference datasets.
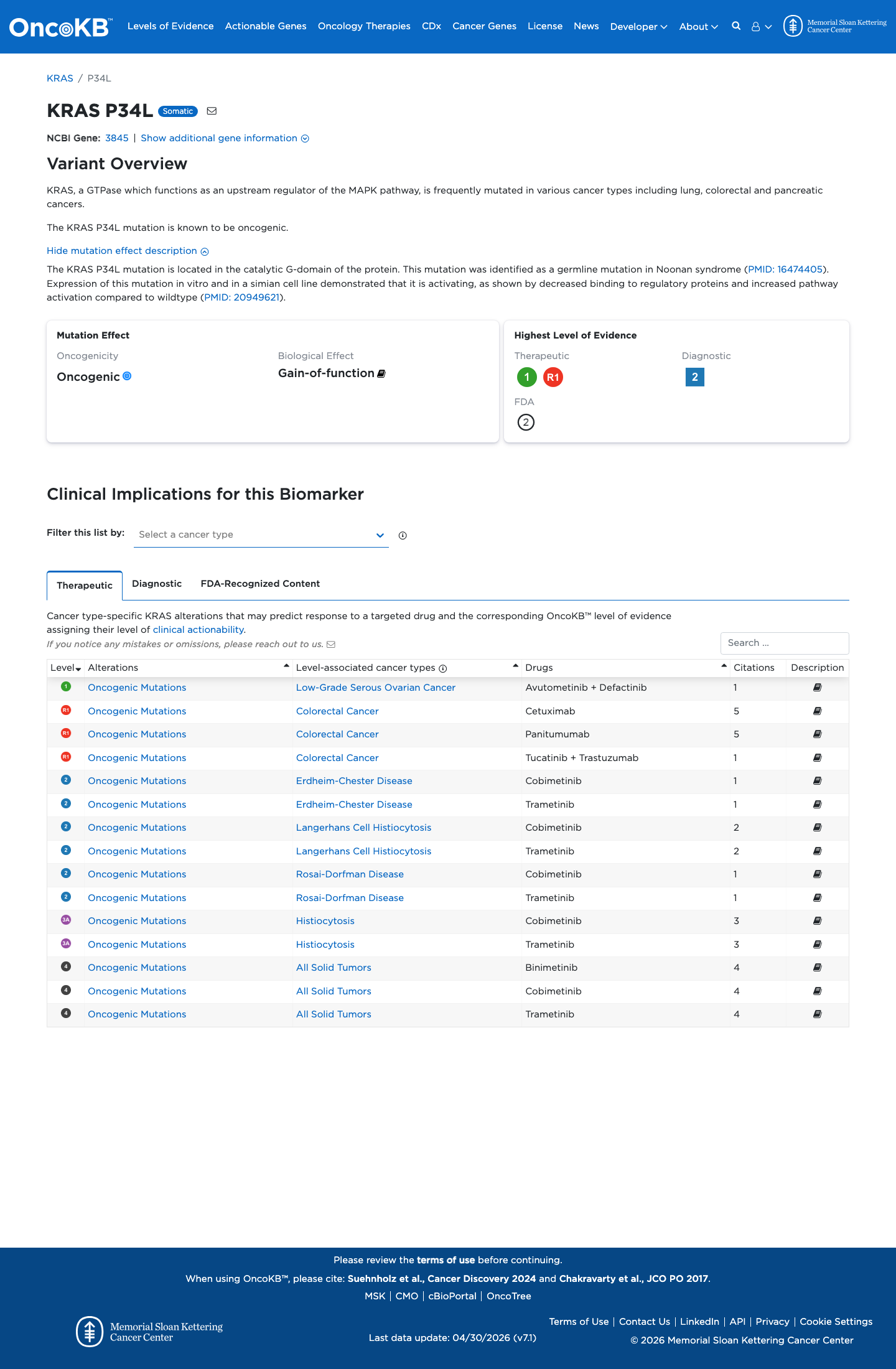
gnomad_v2 ↗ gnomad_v4 ↗In published and VCEP-approved functional studies, this variant showed abnormal KRAS signaling behavior, including increased active GTP-bound KRAS and markedly reduced GAP-stimulated GTP hydrolysis with downstream pathway activation, consistent with a damaging gain-of-function effect.
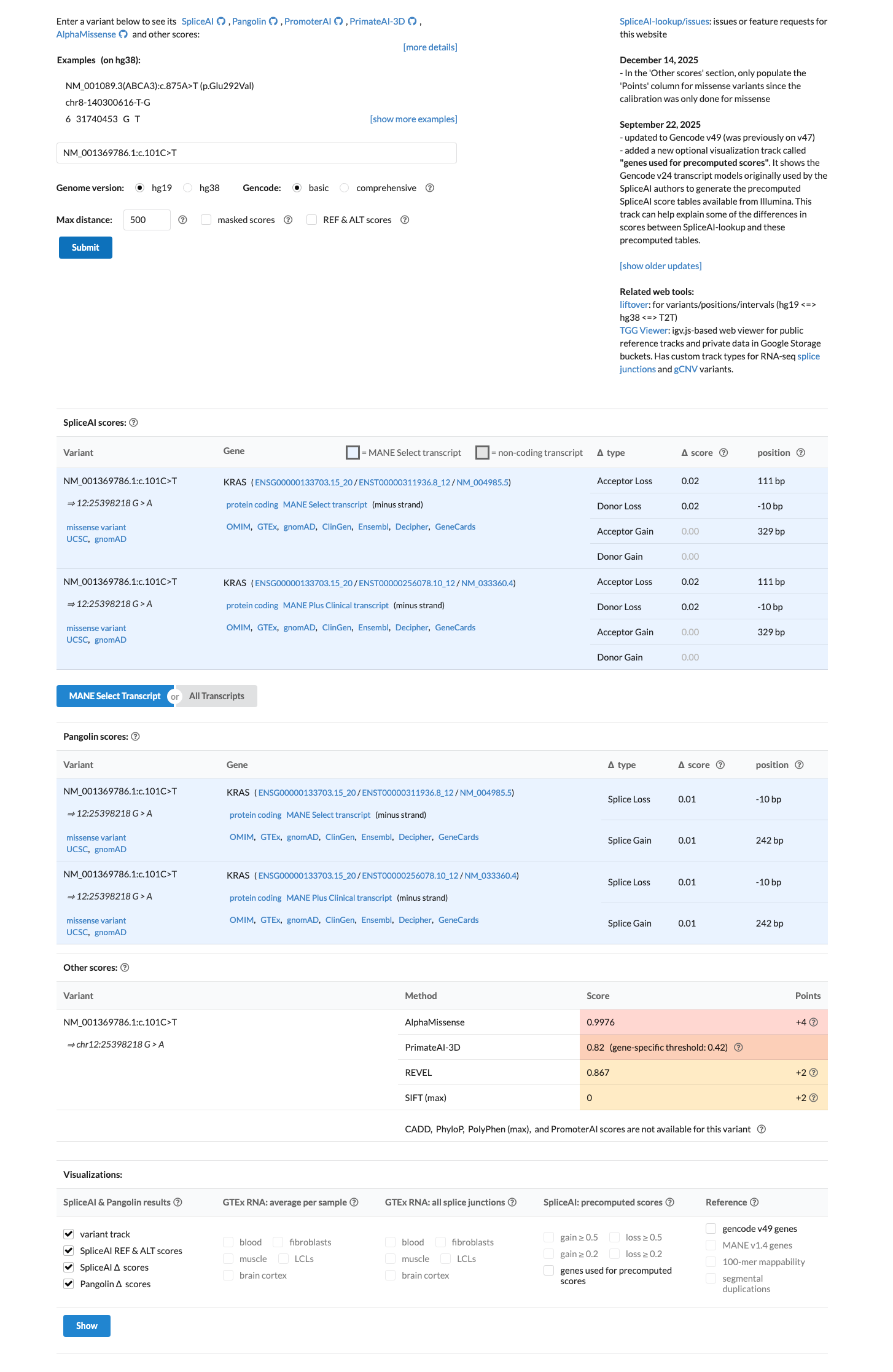
PMID:20949621 ↗Computational evidence supports a deleterious missense effect, with REVEL 0.867 above the KRAS RASopathy PP3 threshold of 0.7, BayesDel 0.425277 supportive of pathogenicity, and SpliceAI showing no meaningful splice disruption (maximum delta score 0.02).
spliceai ↗ cspec ↗