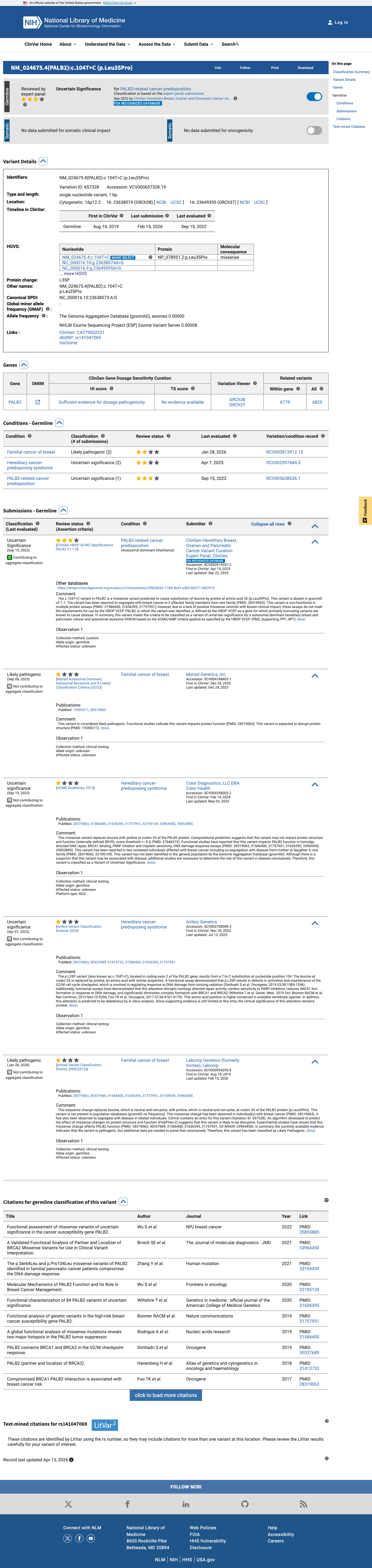
The PALB2 c.104T>C (p.Leu35Pro) variant has not been observed in COSMIC and has been reported in ClinVar with an overall expert-panel classification of uncertain significance, alongside individual clinical laboratory submissions of likely pathogenic and uncertain significance.
clinvar ↗This variant is absent from gnomAD v2.1 and is present only once in gnomAD v4.1 at an overall allele frequency of 0.0000619617% (1/1613900 alleles), which is below the PALB2 PM2_Supporting threshold of 0.000333%.
gnomad_v2 ↗ gnomad_v4 ↗ cspec ↗In published functional studies, p.(Leu35Pro) disrupted the BRCA1-PALB2 interaction, impaired homologous recombination and homology-directed repair, reduced RAD51 foci formation, and increased sensitivity to cisplatin and PARP inhibition, findings consistent with a damaging effect on PALB2 function.
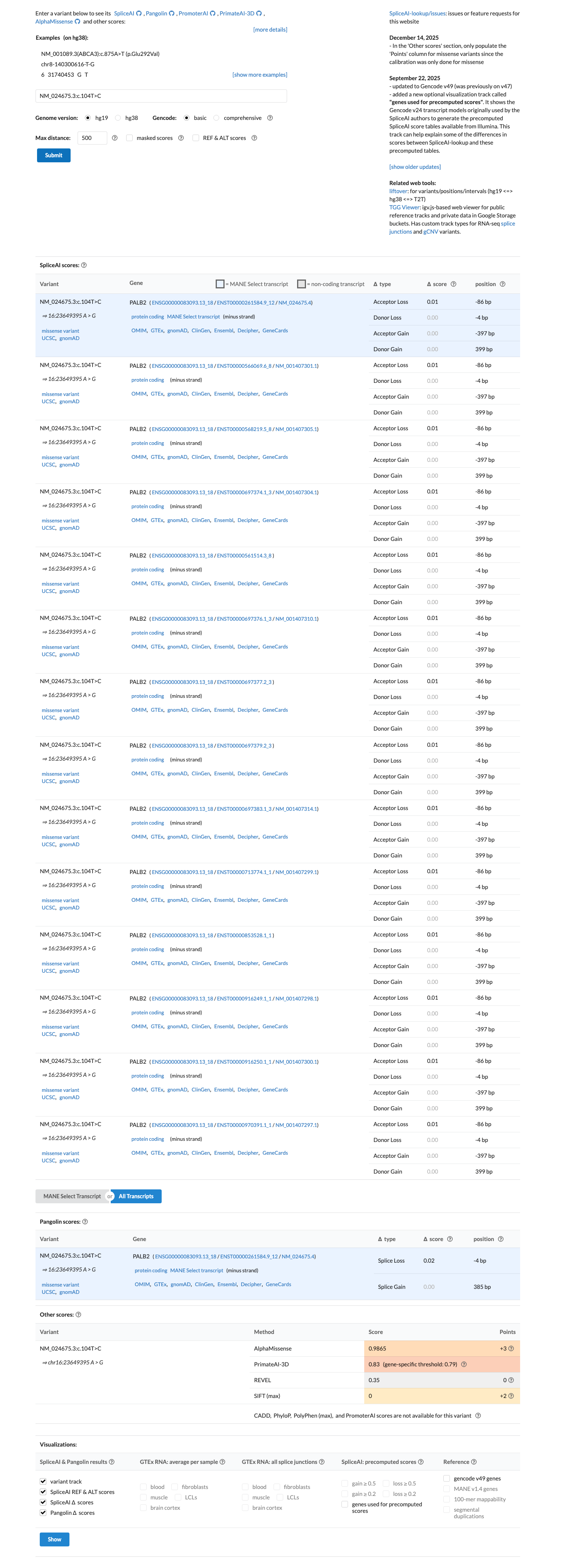
PMID:28319063 ↗ PMID:31636395 ↗In silico splice prediction does not support an RNA effect, with SpliceAI showing a maximum delta score of 0.01; REVEL (0.35) and BayesDel (0.236696) scores are available, but the PALB2 specification does not use missense predictor data for PP3 or BP4.
spliceai ↗ cspec ↗