Classification rationale
1
The BRCA2 c.9117G>A (p.Pro3039=) variant has been reported in ClinVar as Pathogenic with expert panel review.
clinvar ↗2
This variant is rare in population databases, with AF 4.03e-06 (1/248378 alleles) in gnomAD v2.1 and AF 3.72e-06 (6/1613038 alleles) in gnomAD v4.1.
gnomad_v2 ↗ gnomad_v4 ↗3
In a published RNA study, this variant caused exon 23 skipping in puromycin-treated lymphocytes, supporting an abnormal splicing effect consistent with loss of function.
cspec ↗4
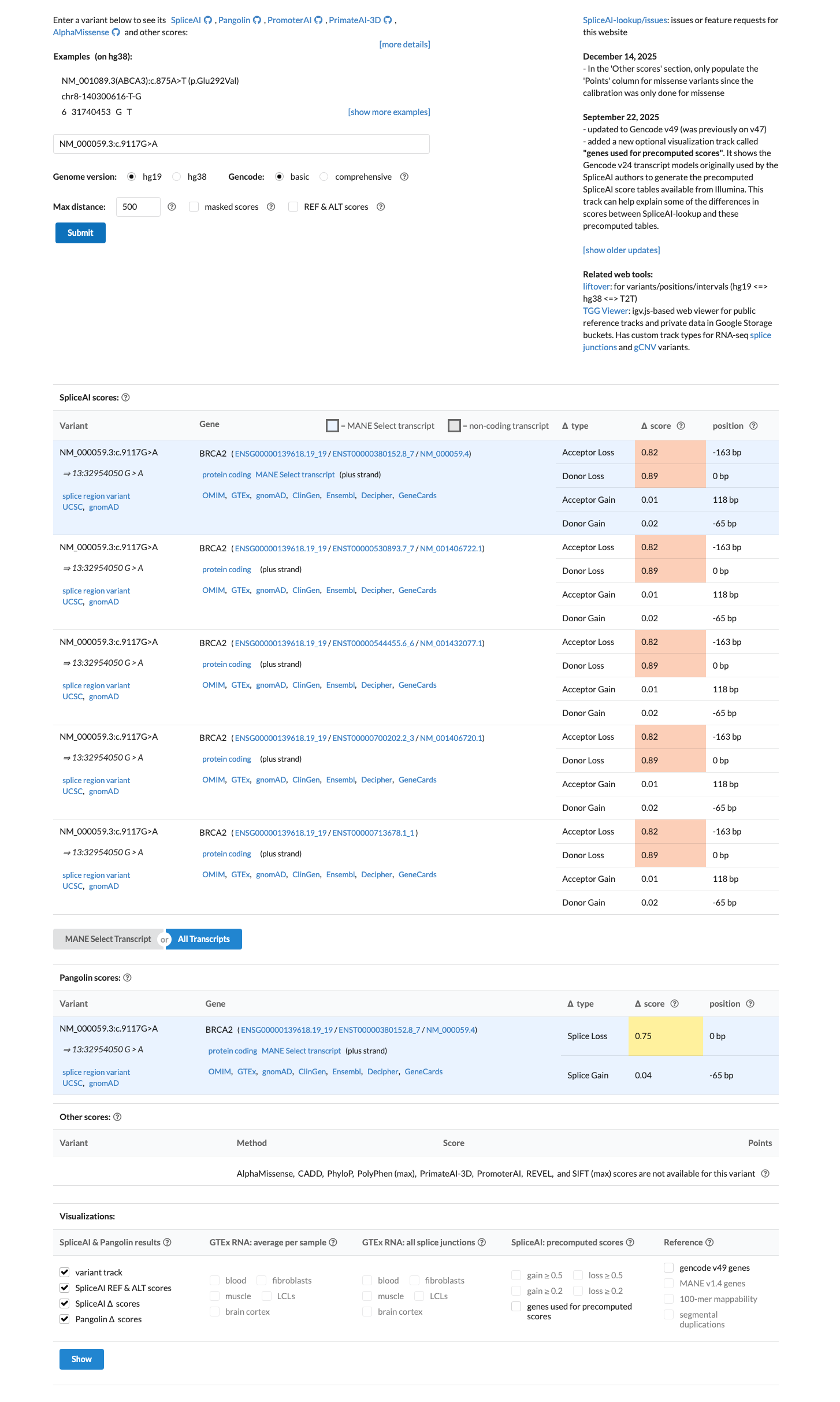
SpliceAI predicts strong splice disruption with a maximum delta score of 0.89, which supports a deleterious splicing effect.
spliceai ↗ cspec ↗5
In the BRCA2-specific clinical-history likelihood dataset, this variant had an LR of 3.91 in 12 probands, meeting ENIGMA PP4 at supporting strength.
PMID:31853058 ↗