Classification rationale
1
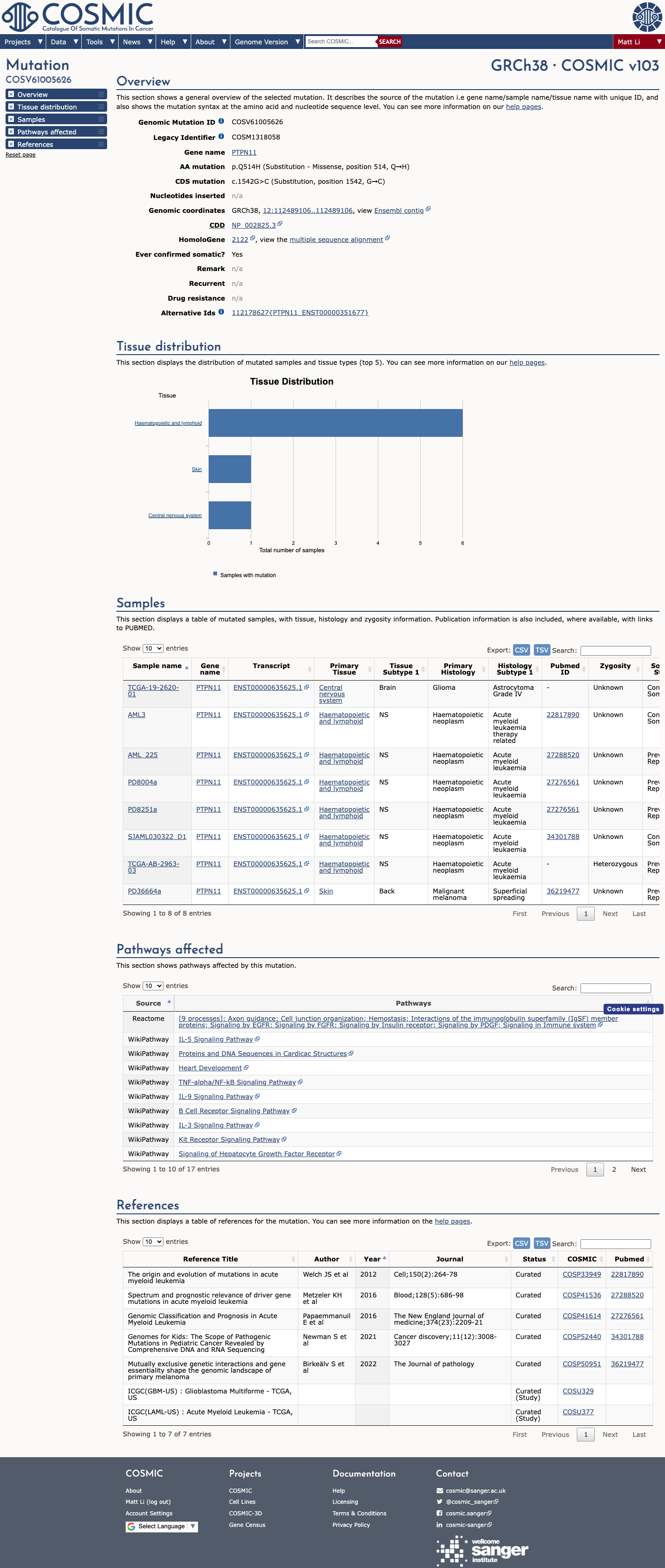
The PTPN11 c.1542G>C (p.Gln514His) variant has been observed in somatic cancers in COSMIC (8 occurrences) and has been reported in ClinVar as pathogenic, including review by the ClinGen RASopathy Variant Curation Expert Panel.
clinvar ↗2
This variant is absent from gnomAD v2.1 and absent from gnomAD v4.1, supporting rarity in population controls.
gnomad_v2 ↗ gnomad_v4 ↗3
ClinVar also documents a different nucleotide change producing the same amino acid substitution, supporting PS1, and in silico results support a deleterious missense effect with REVEL 0.94 above the RASopathy VCEP PP3 threshold of 0.7, BayesDel 0.563665, and no predicted splice impact by SpliceAI (max delta score 0.00).
clinvar ↗ cspec ↗ spliceai ↗