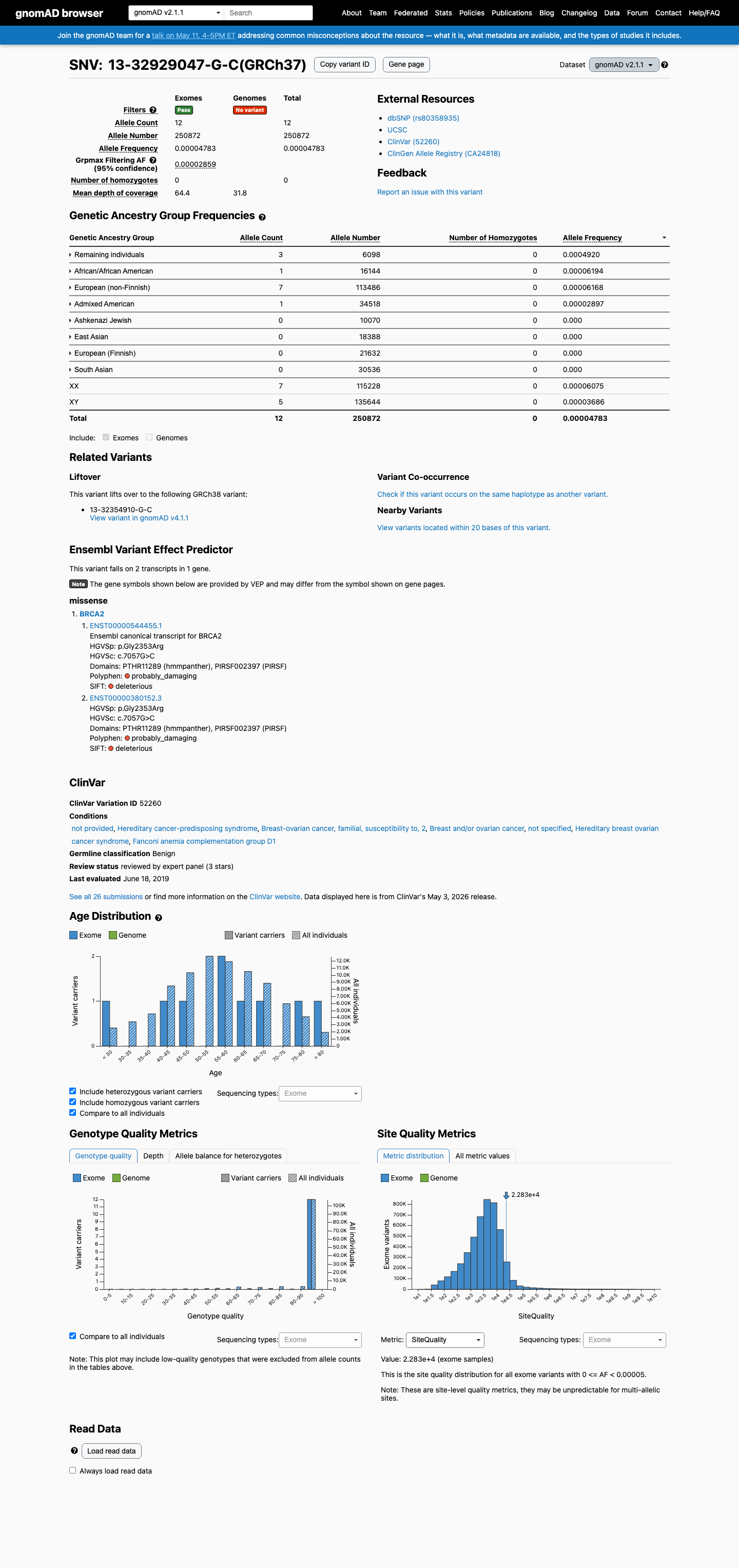
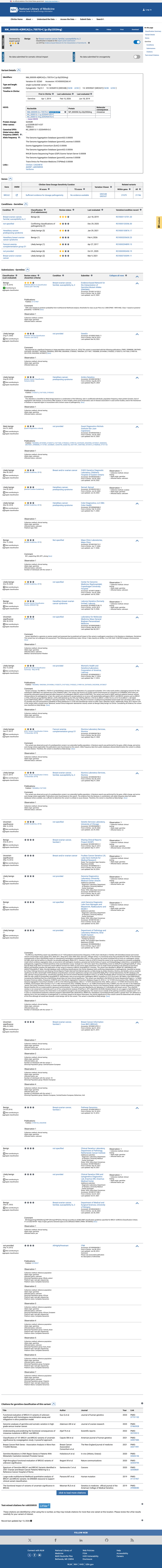
The BRCA2 c.7057G>C (p.Gly2353Arg) variant has been reported in ClinVar, where the aggregate record includes an ENIGMA expert panel Benign classification.
clinvar ↗This variant is present in gnomAD, with grpmax filter allele frequencies of 2.859e-05 in v2.1 and 8.841e-05 in v4.1, which are above the ENIGMA BS1_Supporting threshold of 0.00002 and do not reach the BS1 Strong threshold of 0.0001.
gnomad_v2 ↗ gnomad_v4 ↗ cspec ↗In a calibrated BRCA2 functional study, this variant showed protein function similar to benign control variants, including observed complementation and an HDR score of 82, and ENIGMA Table 9 assigns BS3 Strong.
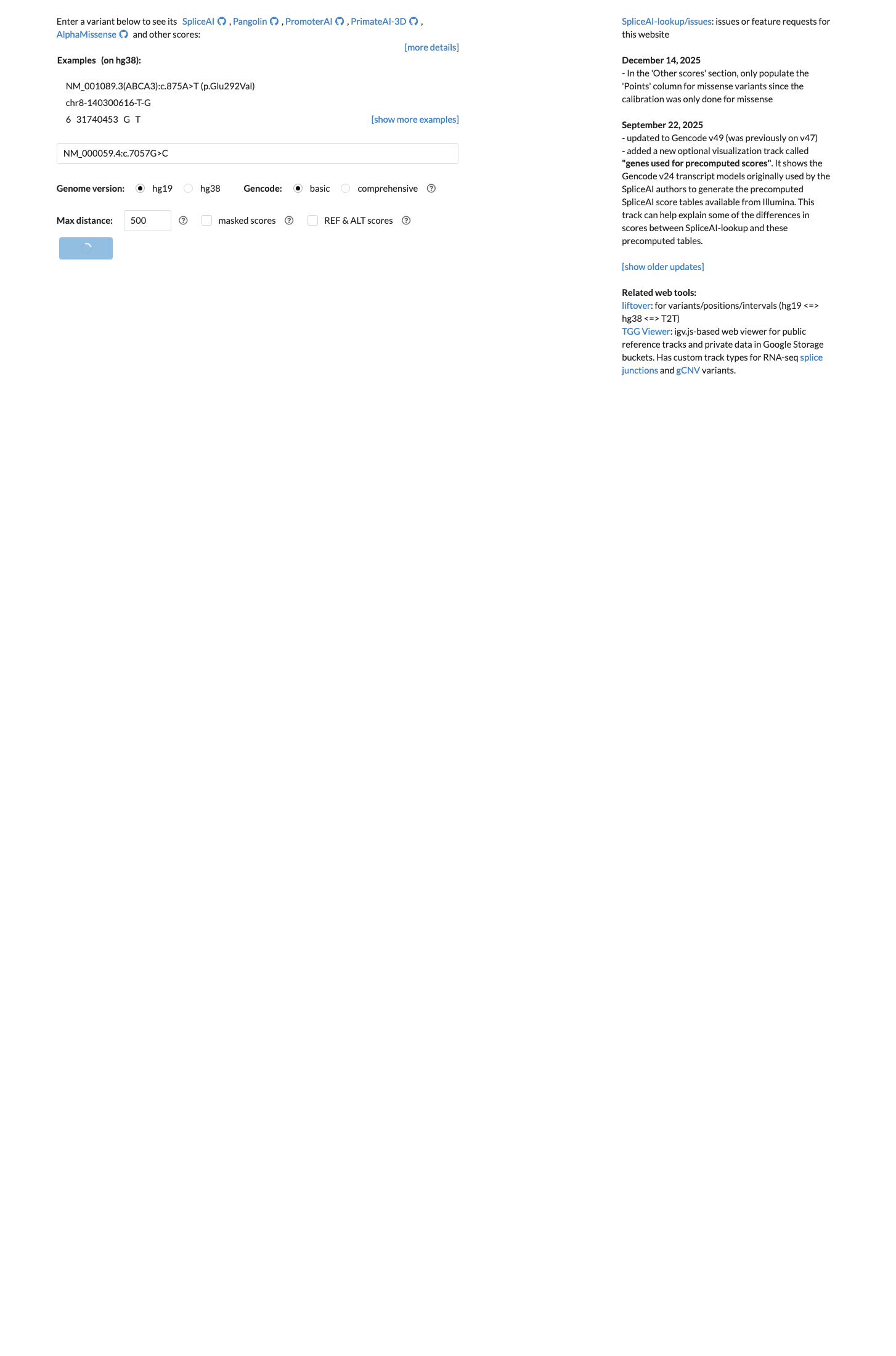
cspec ↗Computational evidence does not support a damaging effect: SpliceAI predicts no significant splice impact with a max delta score of 0.01, BayesDel no-AF is -0.110167, and codon 2353 lies outside the BRCA2 ENIGMA clinically important missense domains, supporting BP1 rather than PP3.
spliceai ↗ cspec ↗