The PALB2 c.682C>T (p.Gln228Ter; p.Q228*) variant has been reported in ClinVar as pathogenic, including review by the ClinGen Hereditary Breast, Ovarian and Pancreatic Cancer Variant Curation Expert Panel.
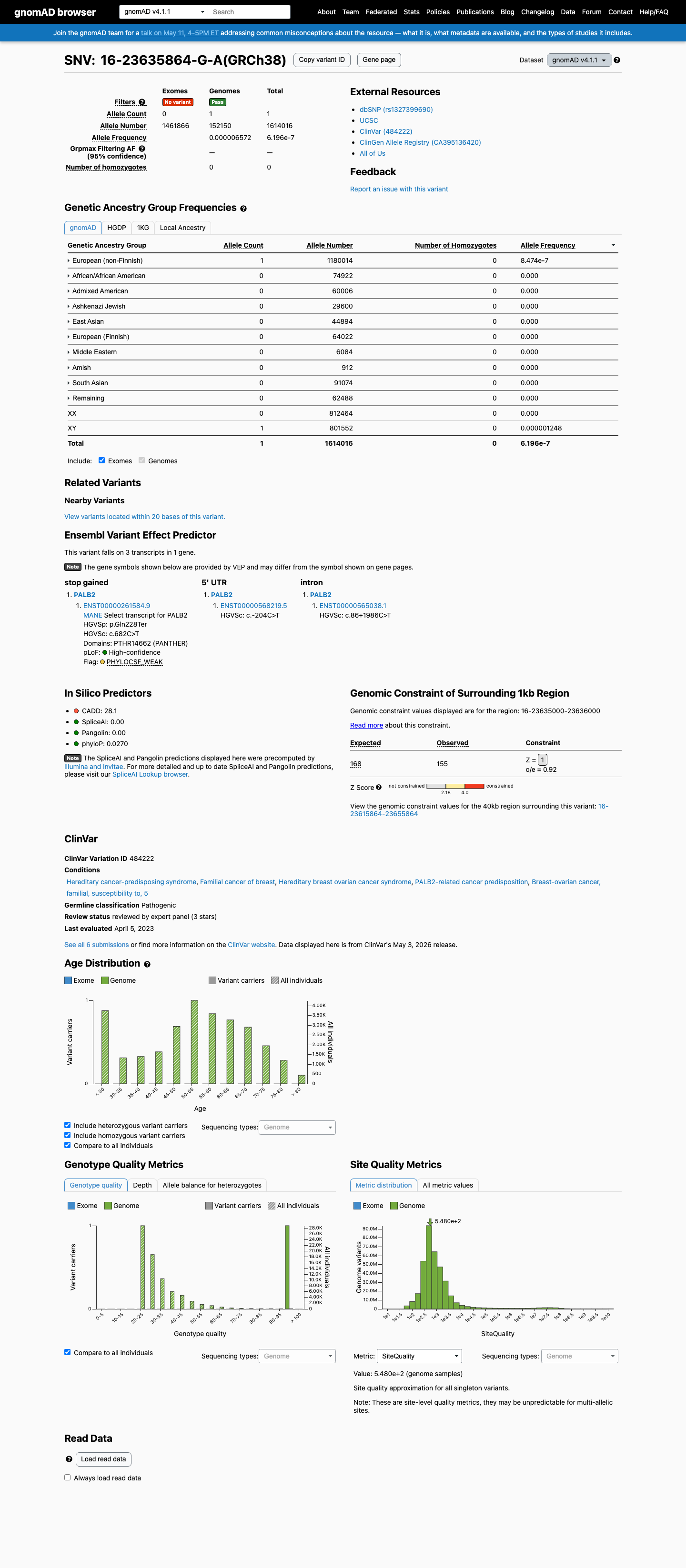
clinvar ↗This variant is absent from gnomAD v2.1 and is present only once in gnomAD v4.1 at an overall allele frequency of 0.00006% (1/1,614,016 alleles), which is below the PALB2 PM2_Supporting threshold of 0.000333%.
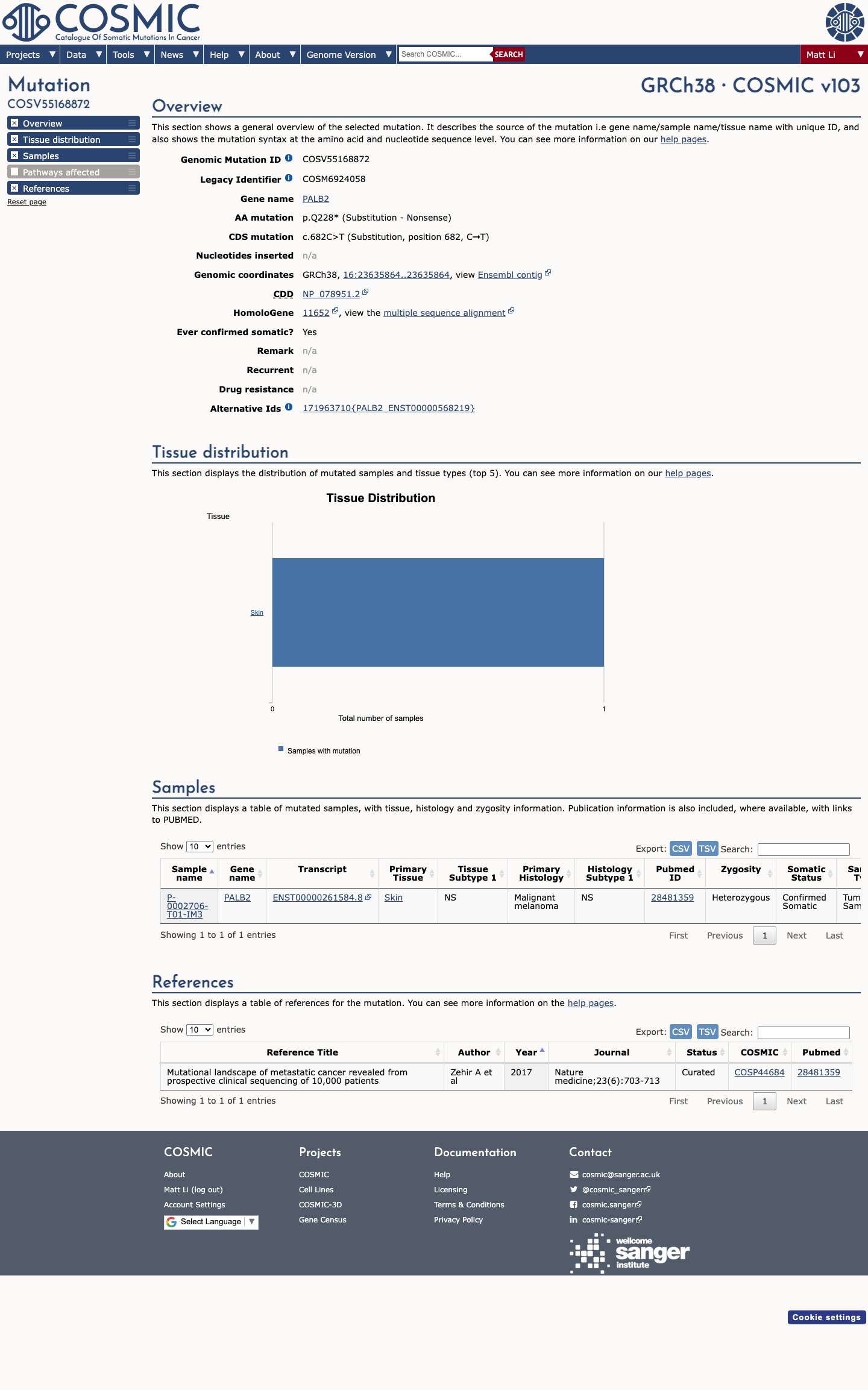
gnomad_v2 ↗ gnomad_v4 ↗ cspec ↗This variant introduces a premature stop codon early in PALB2, and published PALB2 studies together with the PALB2 specification support loss of function as an established disease mechanism for hereditary cancer predisposition.
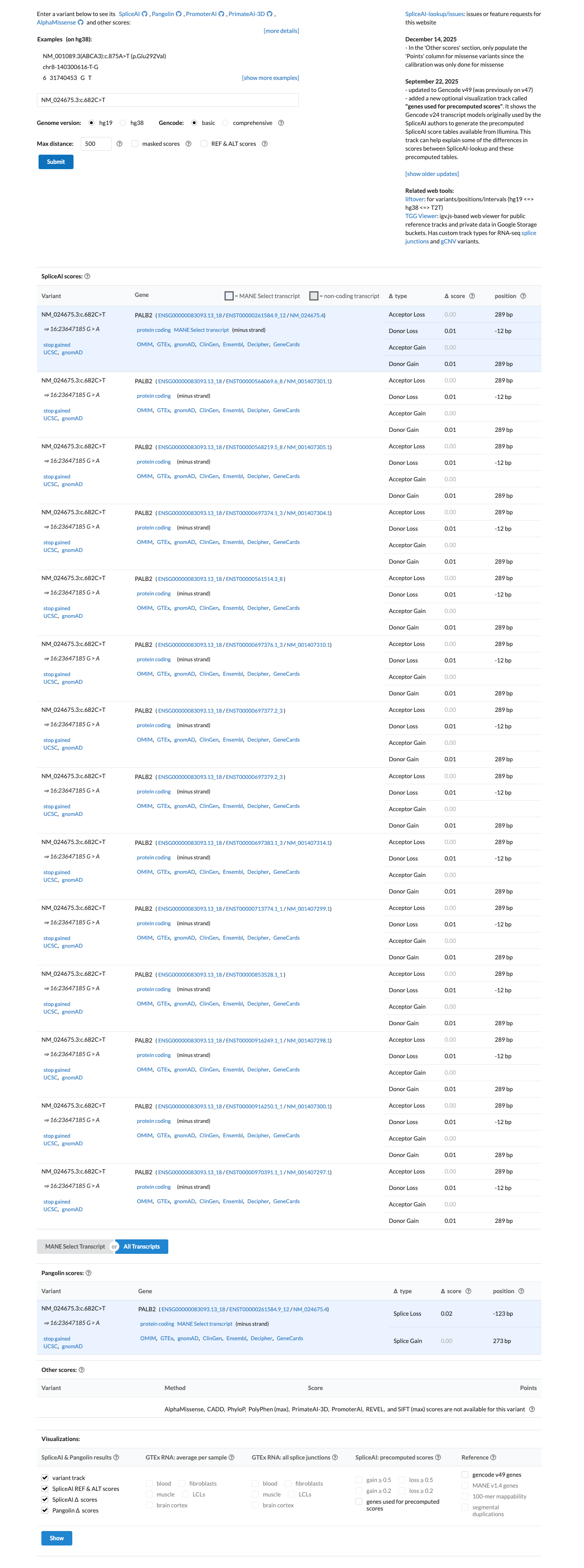
cspec ↗ PMID:17200671 ↗ PMID:25099575 ↗ PMID:28779002 ↗SpliceAI predicts no significant splice impact for this variant (max delta score 0.01), which does not support PP3 and is consistent with the primary consequence being protein truncation rather than altered splicing.
spliceai ↗ cspec ↗