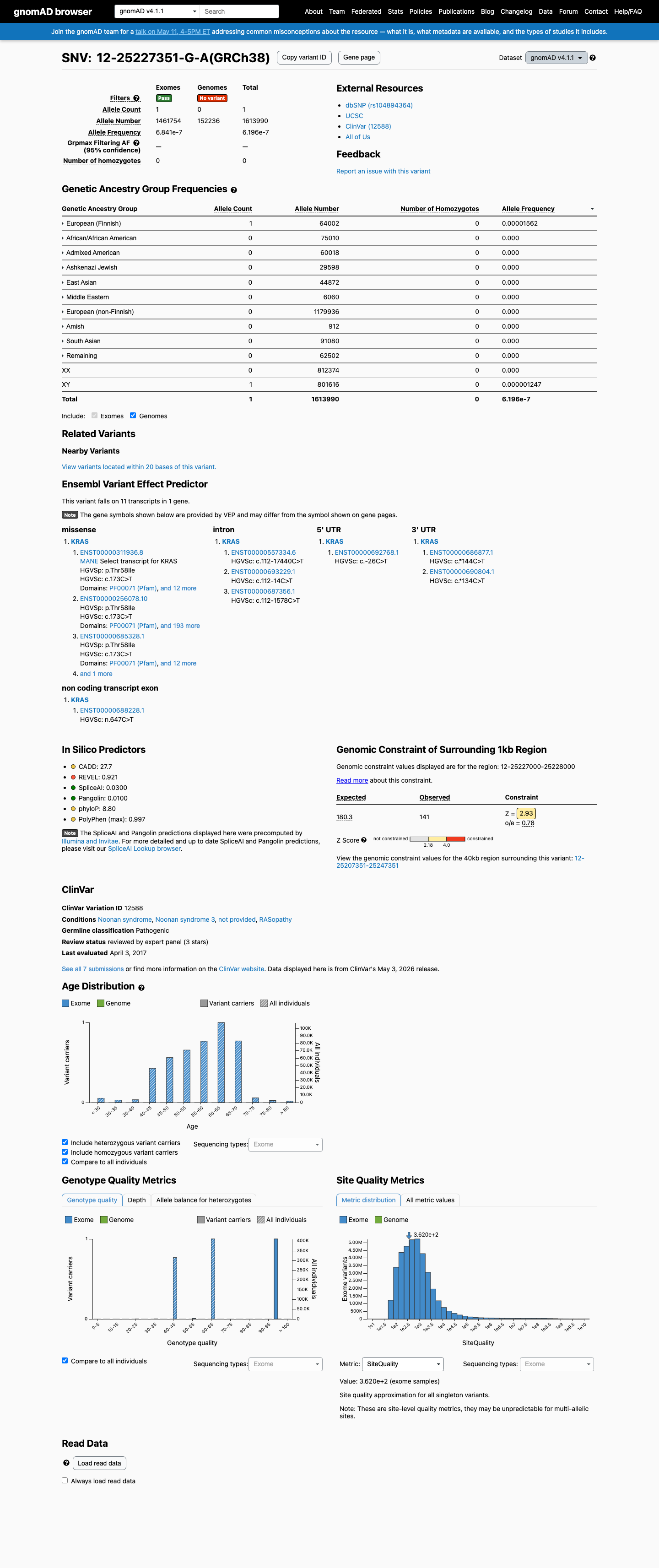
The KRAS c.173C>T (p.Thr58Ile, T58I) variant has been observed in somatic cancer literature and has also been reported in ClinVar as pathogenic, including expert panel review by the ClinGen RASopathy Variant Curation Expert Panel.
PMID:18509354 ↗ clinvar ↗This variant is absent from gnomAD v2.1 and is present only once in gnomAD v4.1 at 1/1613990 alleles (AF 6.19583e-07; 0.00006%), which is far below the KRAS RASopathy VCEP benign frequency thresholds.
gnomad_v2 ↗ gnomad_v4 ↗ cspec ↗In published functional studies, p.Thr58Ile showed defective intrinsic GTP hydrolysis, impaired responsiveness to GAP-mediated regulation, and abnormal downstream signaling; the RASopathy VCEP approved functional study set also includes this variant as a pathogenic or likely pathogenic control in multiple approved assay classes, supporting a gain-of-function effect.
PMID:16474405 ↗ PMID:20949621 ↗Computational evidence supports a deleterious missense effect, with REVEL 0.921 above the VCEP PP3 threshold of 0.7, BayesDel 0.42458, and SpliceAI showing no significant splice effect (max delta score 0.03).
cspec ↗ spliceai ↗