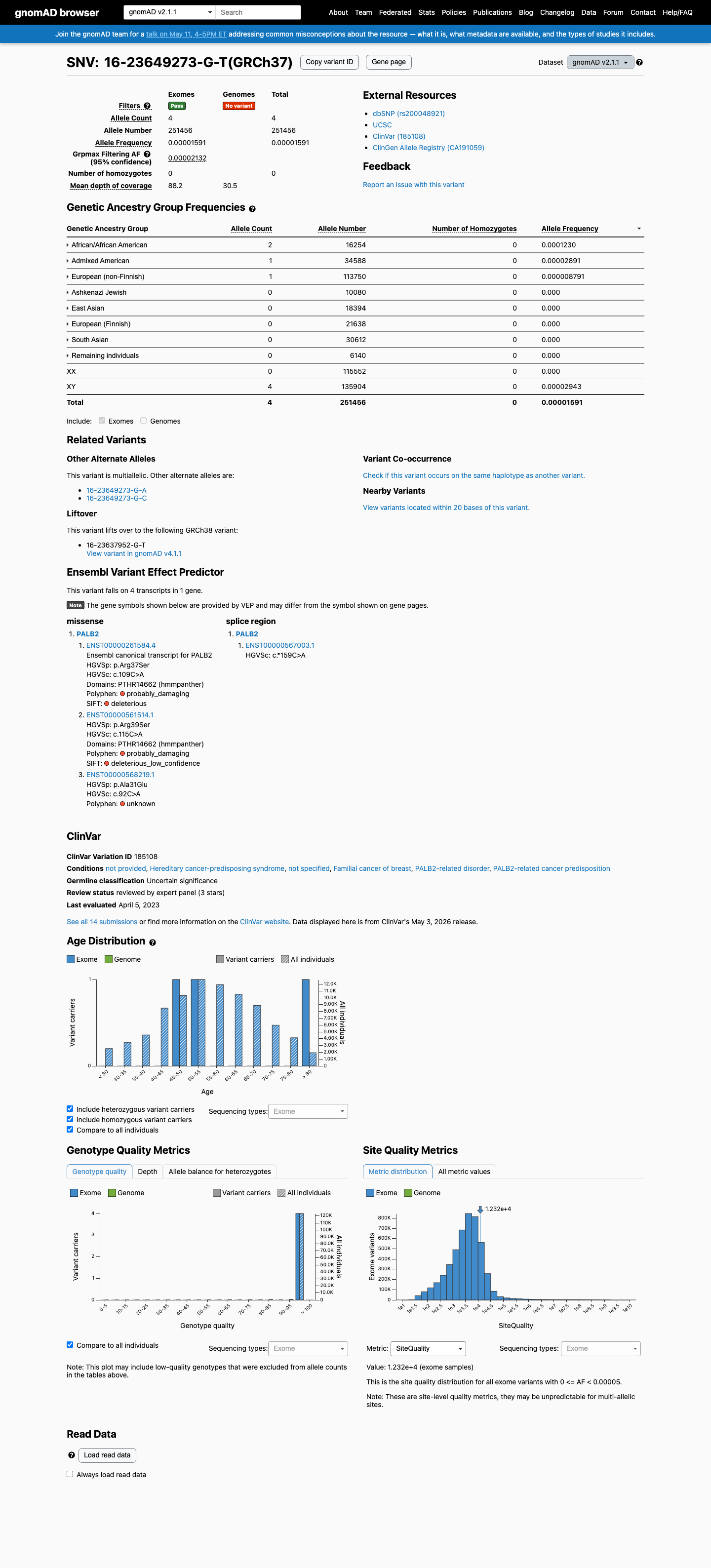
The PALB2 c.109C>A (p.Arg37Ser; p.R37S) variant has been reported in ClinVar with an expert-panel classification of uncertain significance, alongside additional clinical laboratory submissions of uncertain significance and likely benign.
clinvar ↗This variant is present in gnomAD v4.1 at 16/1,612,736 alleles (AF 0.00099%), with a highest observed African/African American subpopulation frequency of 0.01603%, which is above the PALB2 BS1 threshold of 0.01% and below the BA1 threshold of 0.1%.
gnomad_v4 ↗ cspec ↗SpliceAI predicts no meaningful splice effect for this variant (max delta score 0.00), which does not meet the PALB2 PP3 splicing threshold of 0.2 and does not support RNA-based PVS1 application.
spliceai ↗ cspec ↗REVEL (0.195) and BayesDel (0.120598) were noted, but the PALB2 VCEP does not use missense protein predictors for PP3 or BP4, while BP1 is applied to PALB2 missense variants under this framework.
cspec ↗