Classification rationale
1
2
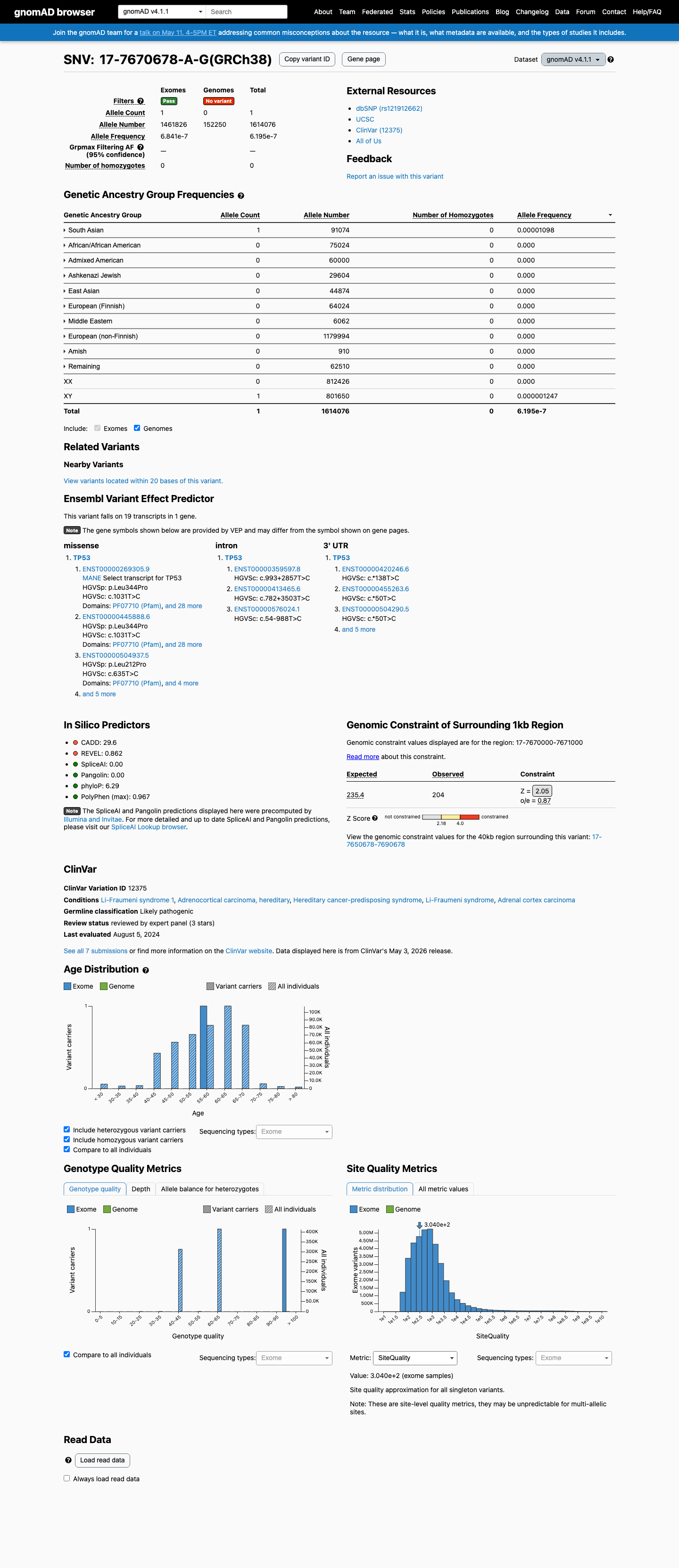
This variant is absent from gnomAD v2.1 and present only once in gnomAD v4.1 (1/1614076 alleles; AF 6.1955e-07), with the highest observed population frequency in South Asian individuals of 1/91074 (AF 1.0980e-05), supporting rarity for TP53.
gnomad_v2 ↗ gnomad_v4 ↗3
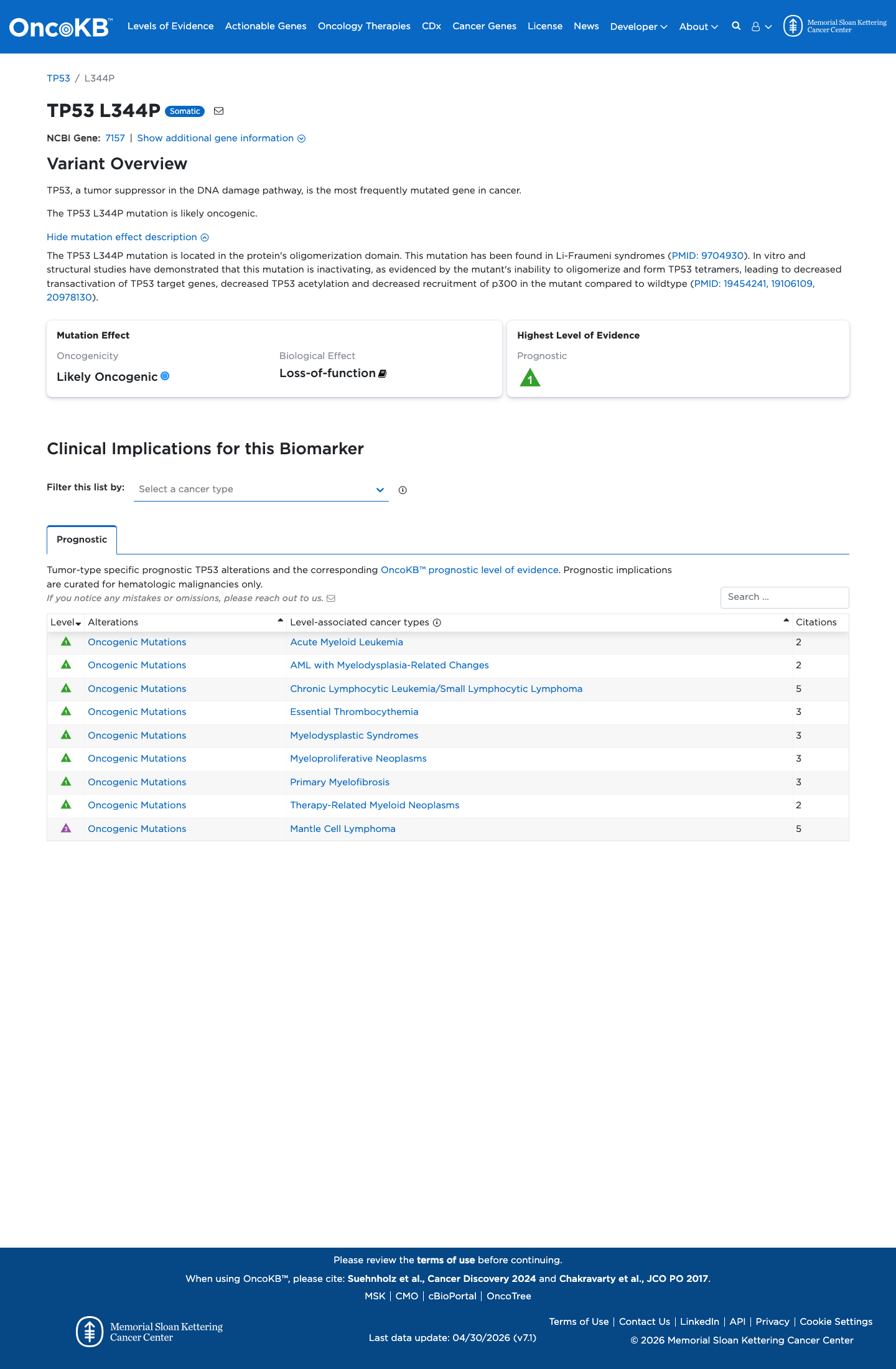
In TP53 functional studies summarized by the TP53 VCEP, p.Leu344Pro showed non-functional activity, loss-of-function behavior across eligible assays, and monomeric oligomerization-domain behavior, supporting a damaging loss-of-function effect.
PMID:12826609 ↗ PMID:9704930 ↗ PMID:19106109 ↗ PMID:19454241 ↗ PMID:20978130 ↗4
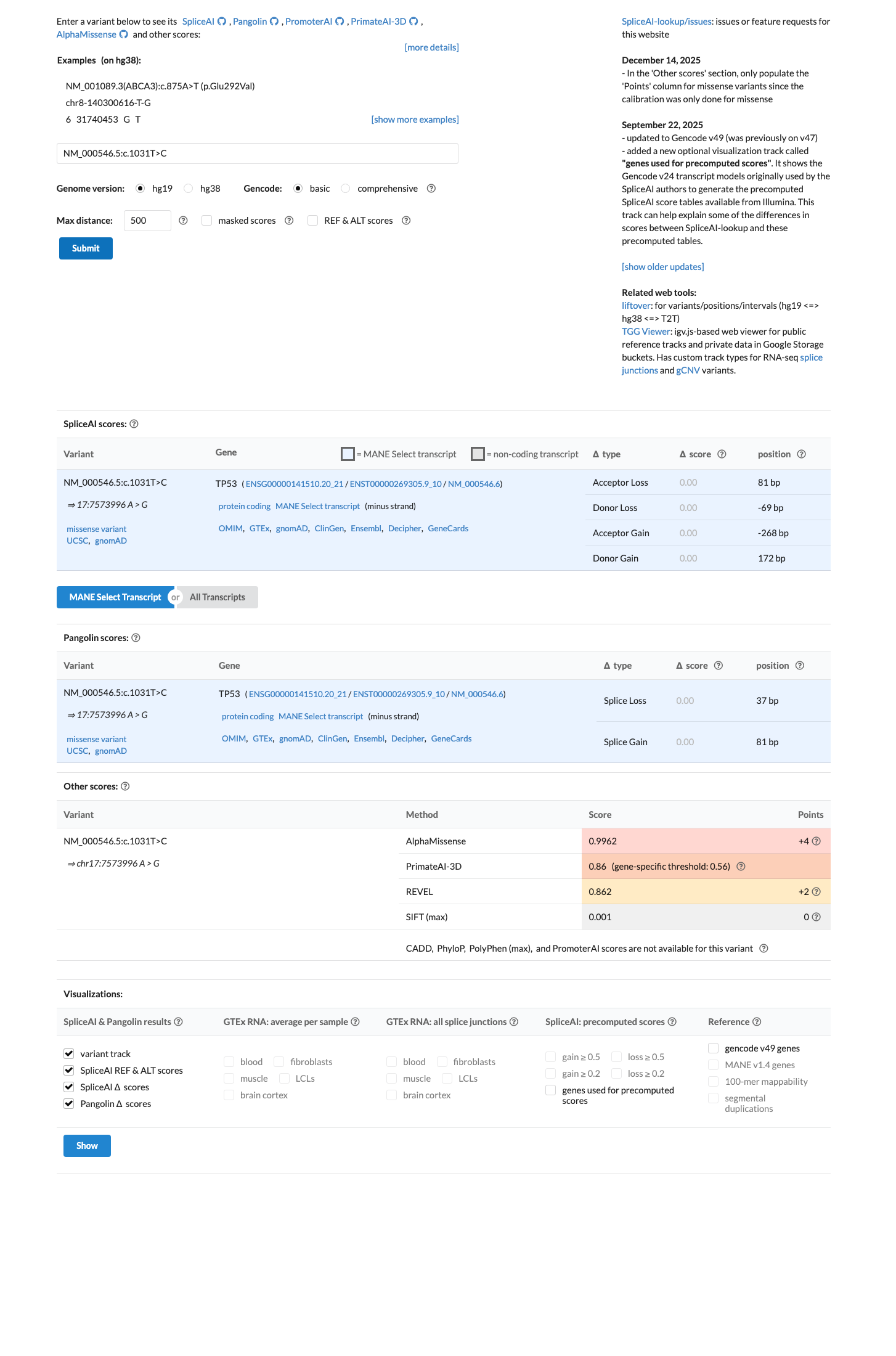
Computational evidence is concordant with a damaging missense effect, with a TP53 VCEP pre-assigned PP3_moderate call, BayesDel 0.485604, and REVEL 0.862.