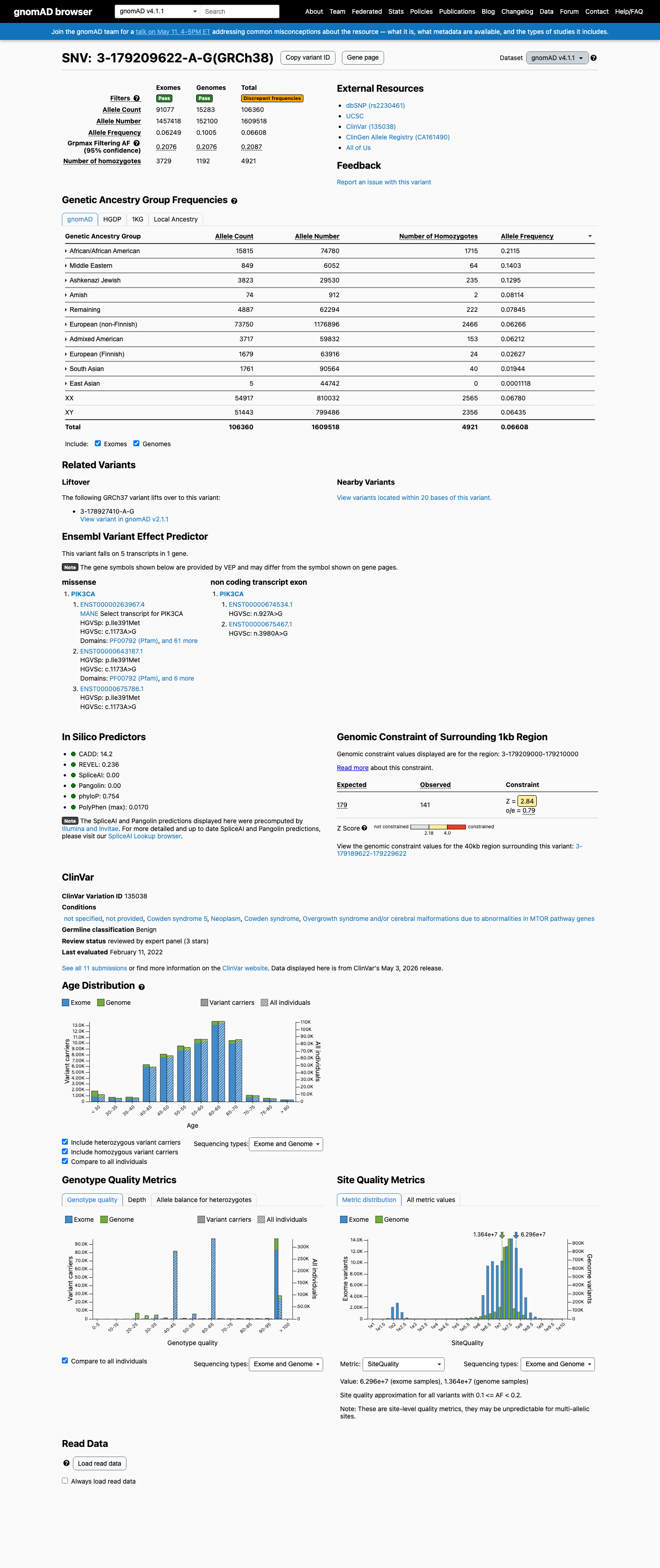
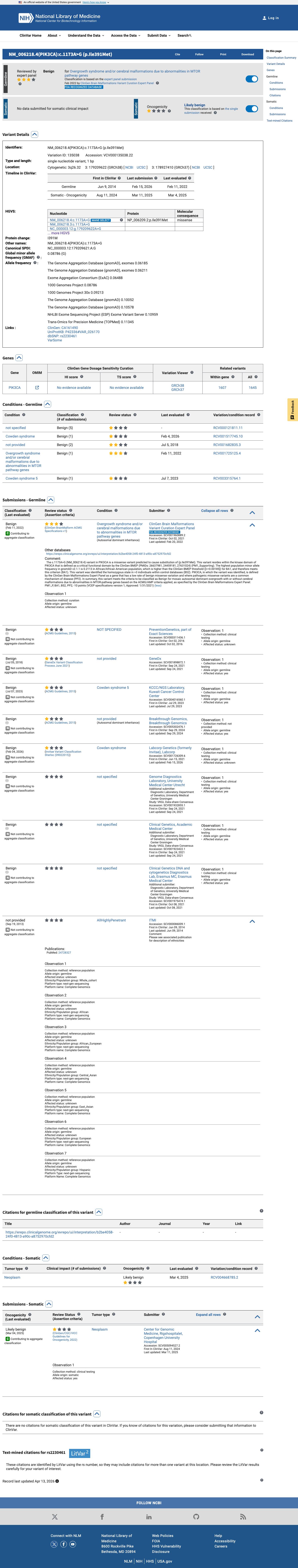
The PIK3CA c.1173A>G (p.Ile391Met) variant has been reported in ClinVar as Benign, including a Benign expert-panel classification from the ClinGen Brain Malformations Variant Curation Expert Panel.
clinvar ↗This variant is common in population databases, with allele frequency 6.57058% in gnomAD v2.1 and 6.60819% in gnomAD v4.1, greatly exceeding the VCEP BA1 threshold of 0.0926% and BS1 threshold of 0.0185%, and it is also observed in numerous homozygous individuals.
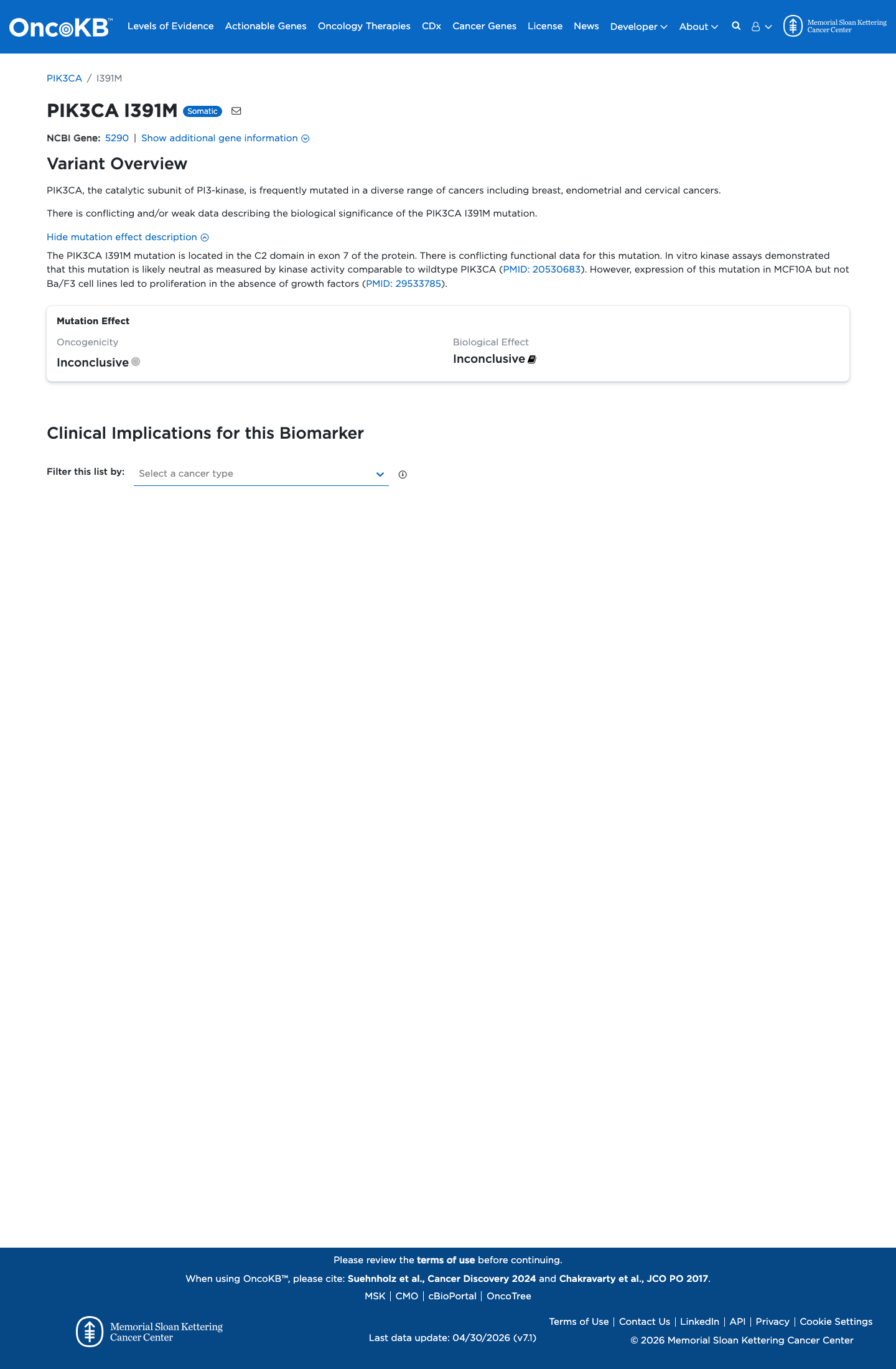
gnomad_v2 ↗ gnomad_v4 ↗ cspec ↗Curated literature resources identified functional publications relevant to this variant, but the retrieved evidence did not provide validated variant-specific assay results sufficient to apply either PS3 or BS3.
oncokb ↗ PMID:20530683 ↗ PMID:29533785 ↗ cspec ↗SpliceAI predicts no significant splice impact for this variant with a maximum delta score of 0.00, while REVEL is 0.236 and BayesDel is -0.391355; however, PP3 is not applicable and BP4 is restricted to non-missense variants in this VCEP framework.
spliceai ↗ cspec ↗