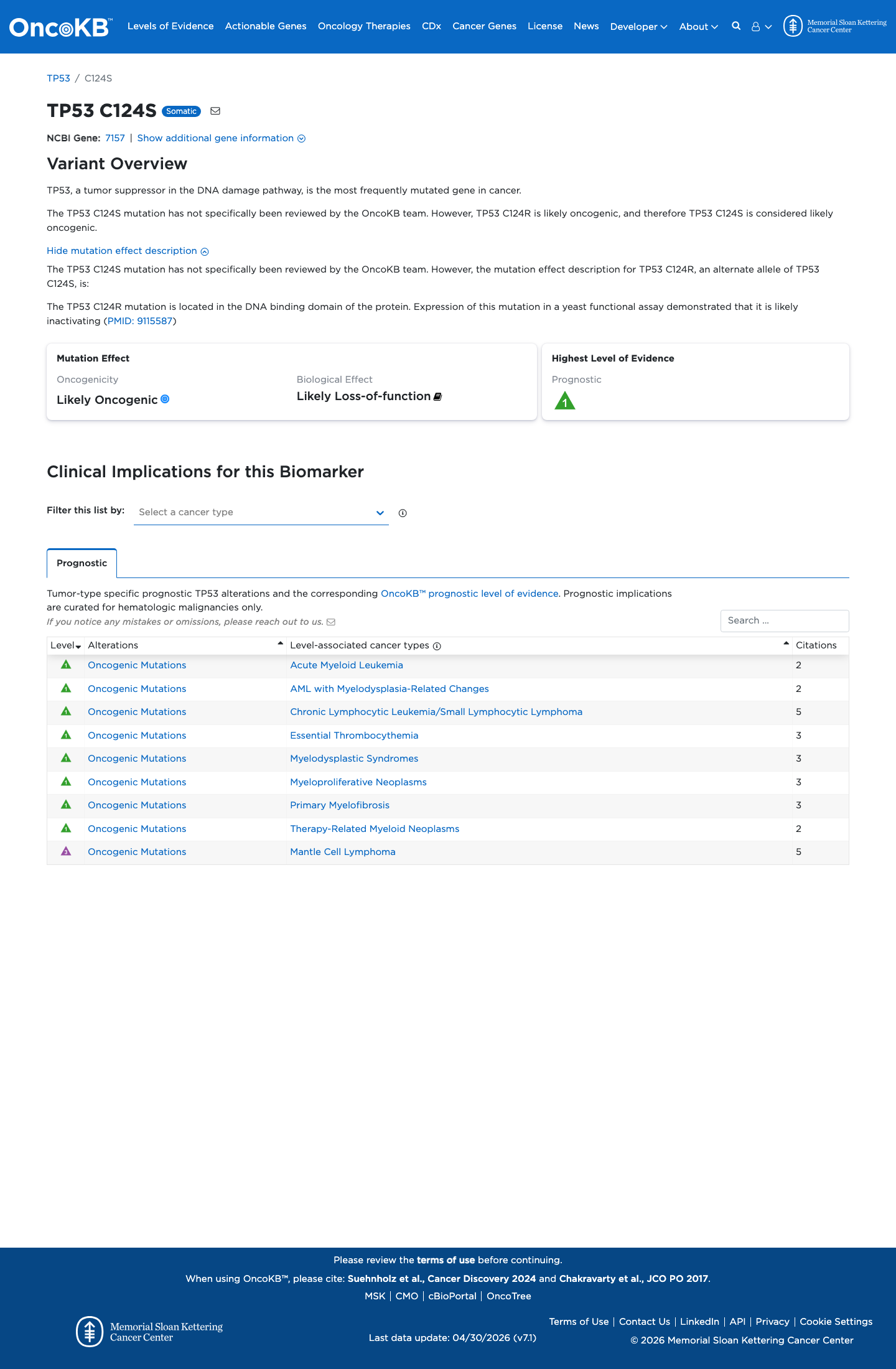
The TP53 c.370T>A (p.Cys124Ser) variant has not been observed in COSMIC and has been reported in ClinVar, where the ClinGen TP53 Variant Curation Expert Panel classified it as Likely Benign.
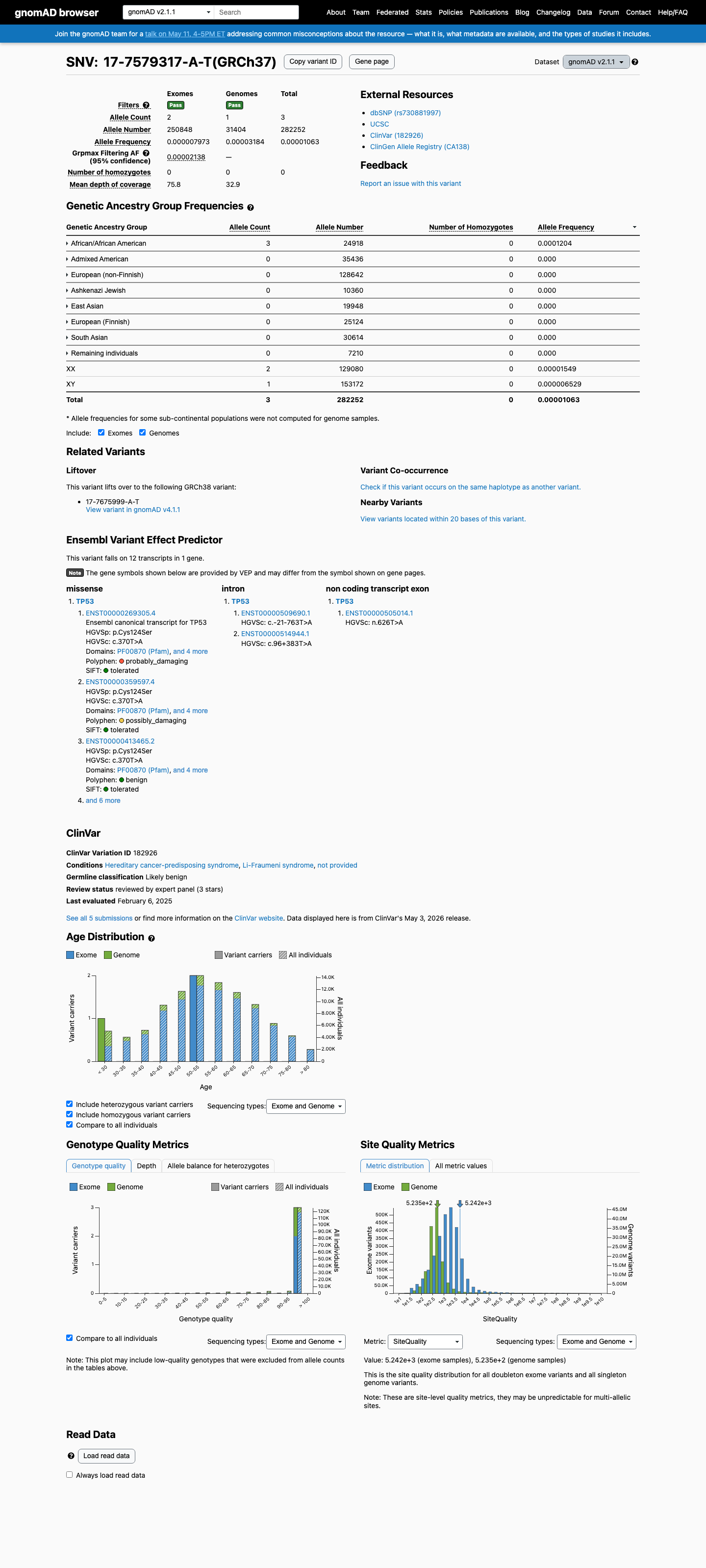
clinvar ↗This variant is present at low frequency in gnomAD, including 3 of 1612186 alleles overall in v4.1 and 3 of 74876 alleles in the African/African American population; the subgroup frequency of 0.0000401 is slightly above the TP53 VCEP PM2 multi-allele threshold of less than 0.00004, so PM2 is not met.
gnomad_v4 ↗ gnomad_v2 ↗ cspec ↗In TP53 VCEP-supported functional datasets, p.Cys124Ser was functional in Kato et al. and showed no loss of function in the Giacomelli and Kotler datasets, supporting BS3.
PMID:12826609 ↗ PMID:29979965 ↗In silico evidence is mixed: the TP53 VCEP bioinformatic worksheet assigns PP3_Moderate to c.370T>A based on Align-GVGD class C65 and BayesDel 0.257897, while SpliceAI is 0.02 and predicts no meaningful splice effect; REVEL is 0.716.
spliceai ↗