Classification rationale
1
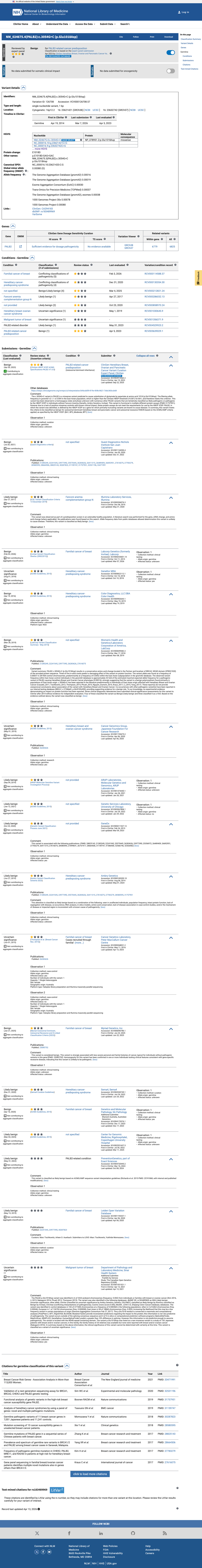
The PALB2 c.3054G>C (p.Glu1018Asp; E1018D) variant has been reported in ClinVar, where the aggregate classification is Benign with expert panel review.
clinvar ↗2
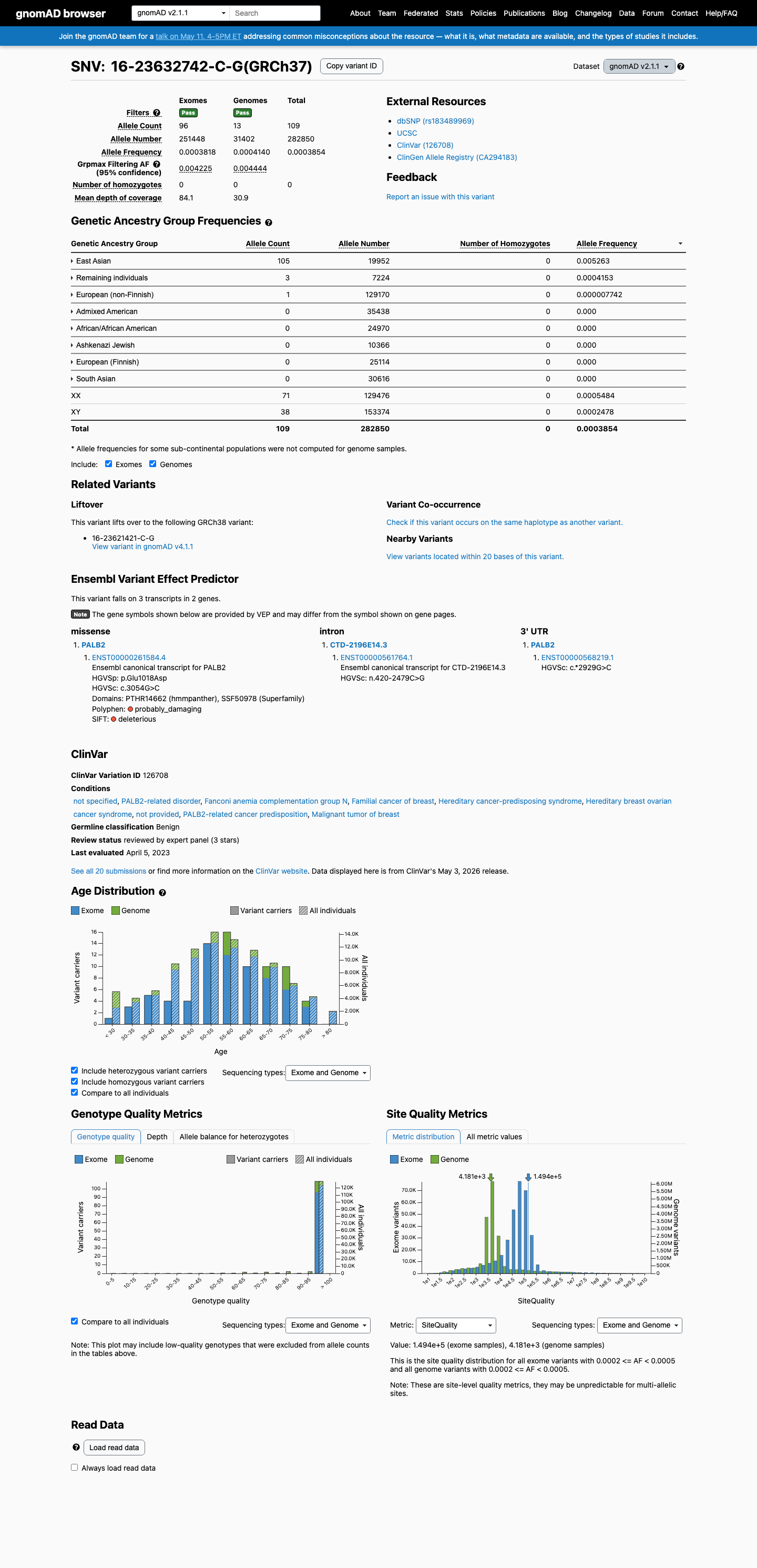
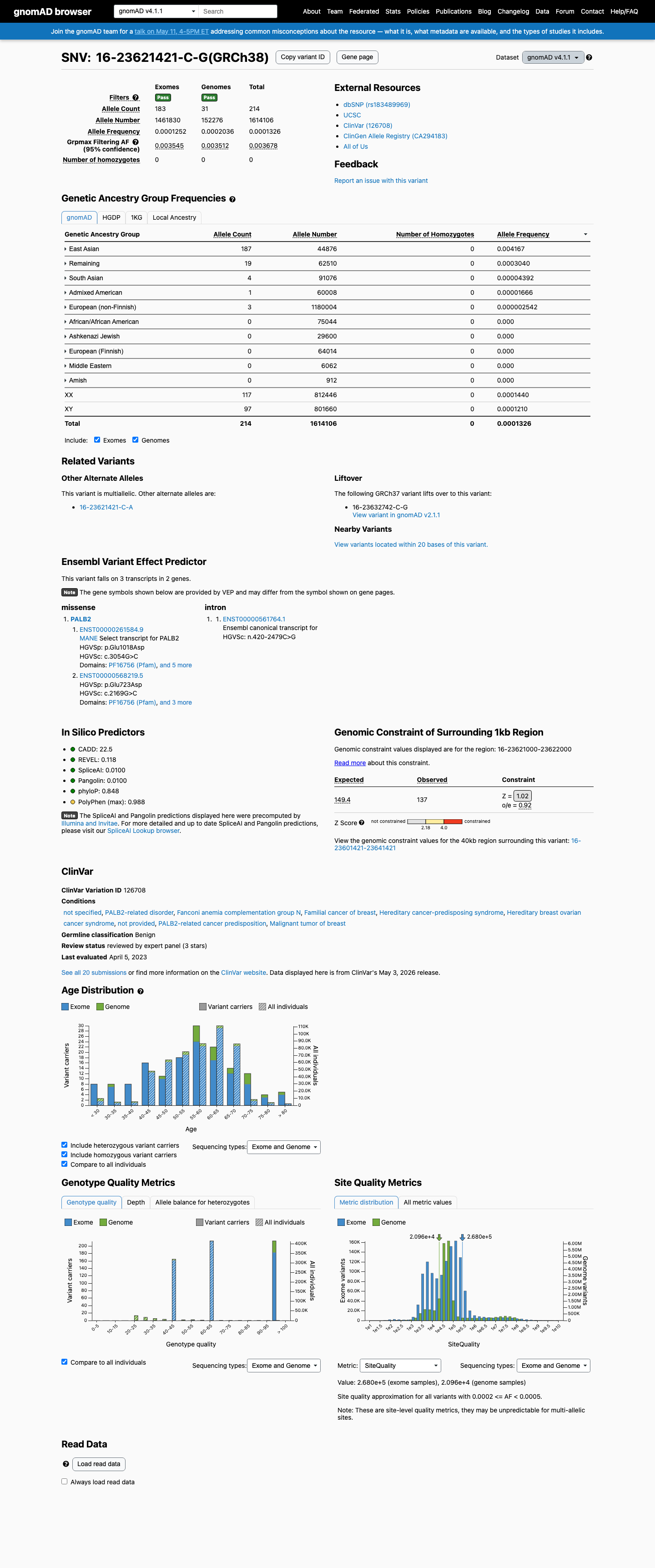
This variant is present in gnomAD v4.1 at 214/1,614,106 alleles (AF 0.01326%), with highest observed East Asian frequency 187/44,876 alleles (AF 0.41670%), which is above the PALB2 BA1 threshold of 0.1%.
gnomad_v4 ↗ cspec ↗3
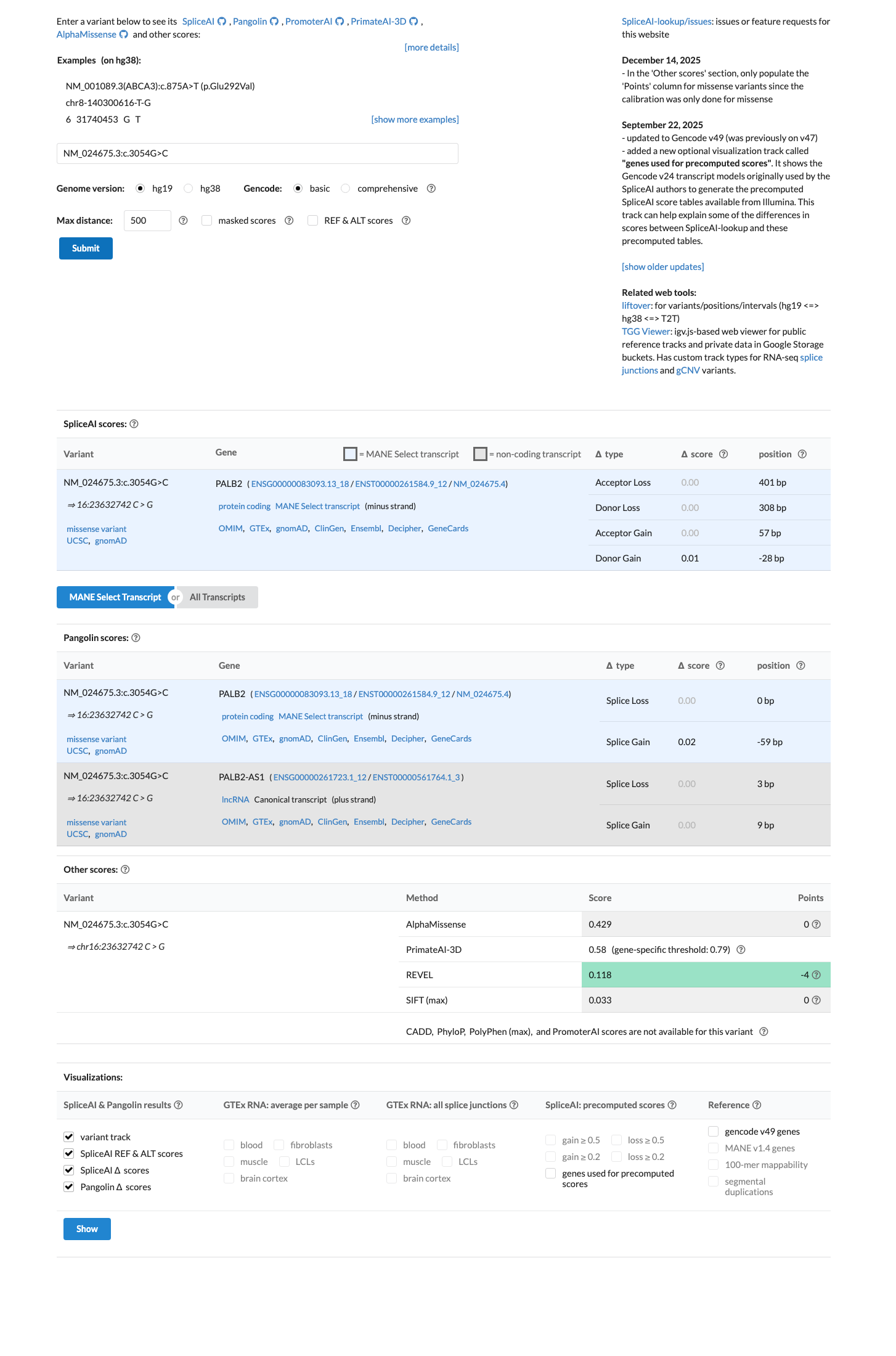
SpliceAI predicts no significant splice effect (max delta score 0.01), below the PALB2 PP3 splice threshold of 0.2; REVEL is 0.118 and BayesDel is -0.42024, and the PALB2 specification does not support missense PP3/BP4 use beyond its stated splice rules.
spliceai ↗ cspec ↗