Classification rationale
1
2
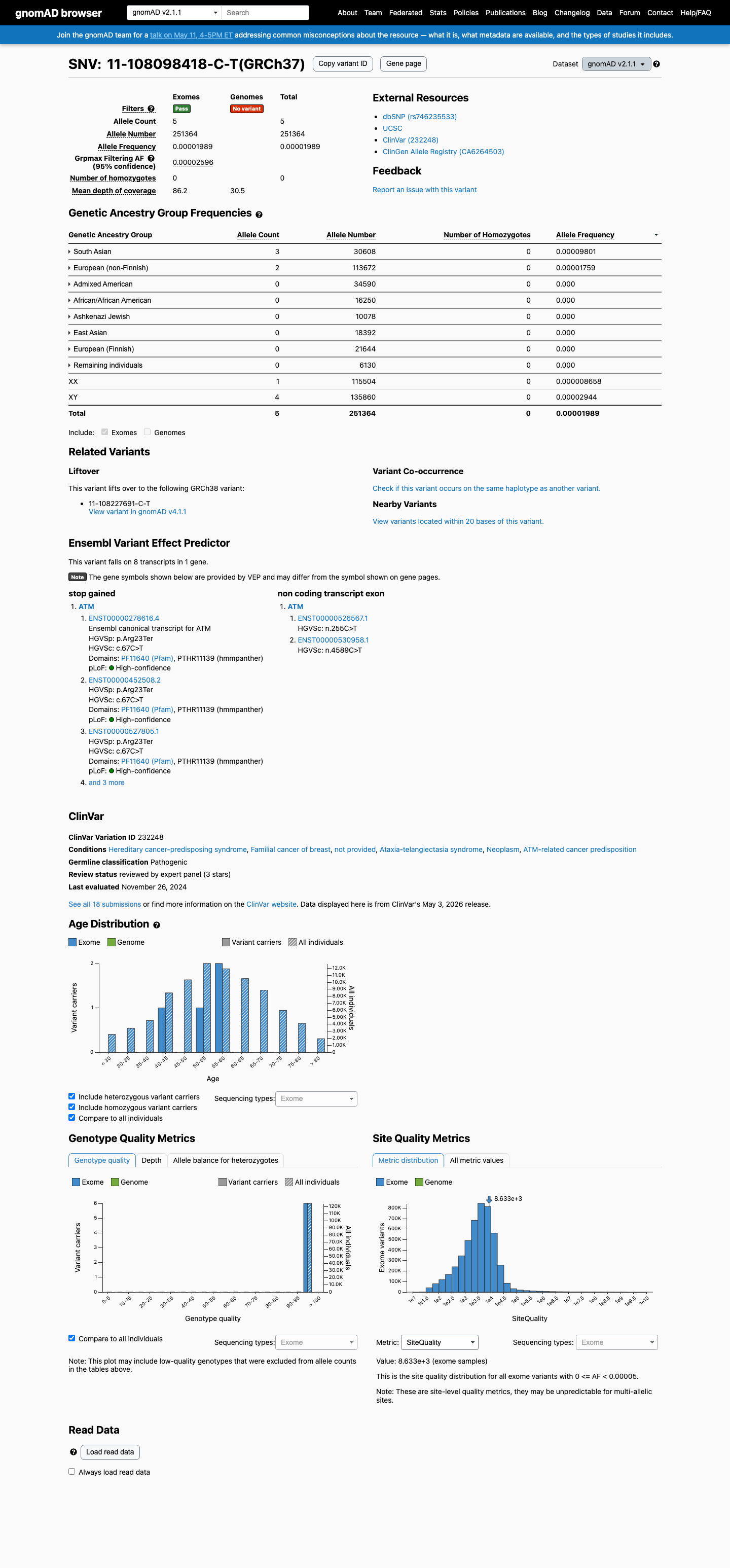
This variant is rare in population databases, with gnomAD v4.1 AF 3.71898e-06 (0.00037%, 6/1,613,346 alleles; no homozygotes), which is below the ATM PM2 threshold of 0.001%.
gnomad_v4 ↗ cspec ↗3
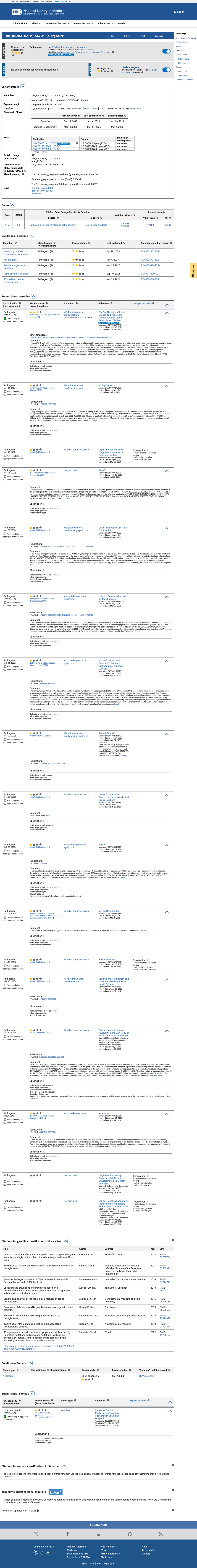
In an ATM supplementary SNV table, this variant was categorized as non-functional with medium-high confidence, which is consistent with a damaging loss-of-function effect, although the available summary alone does not establish ATM VCEP PS3 weighting.
cspec ↗4
This is a nonsense variant predicted to create a very early stop codon; SpliceAI predicts no significant splice impact with a max delta score of 0.00, and the ATM computational PP3/BP4 rules are not used here because they are defined for missense or specified splicing contexts rather than truncating variants.
spliceai ↗ cspec ↗