Classification rationale
1
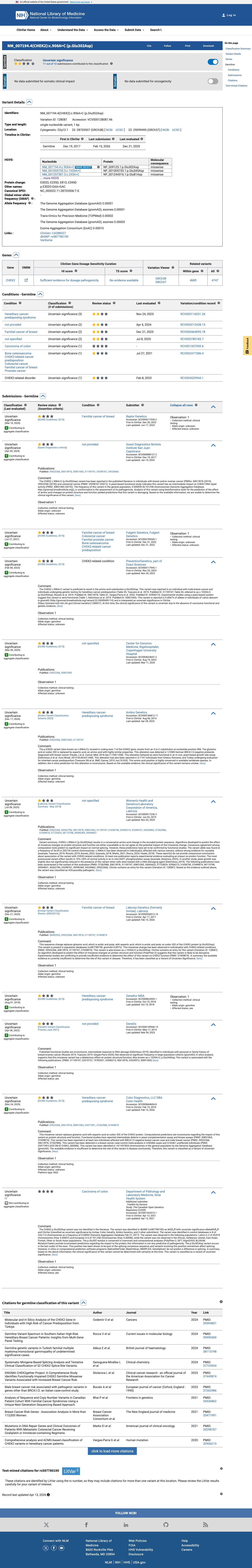
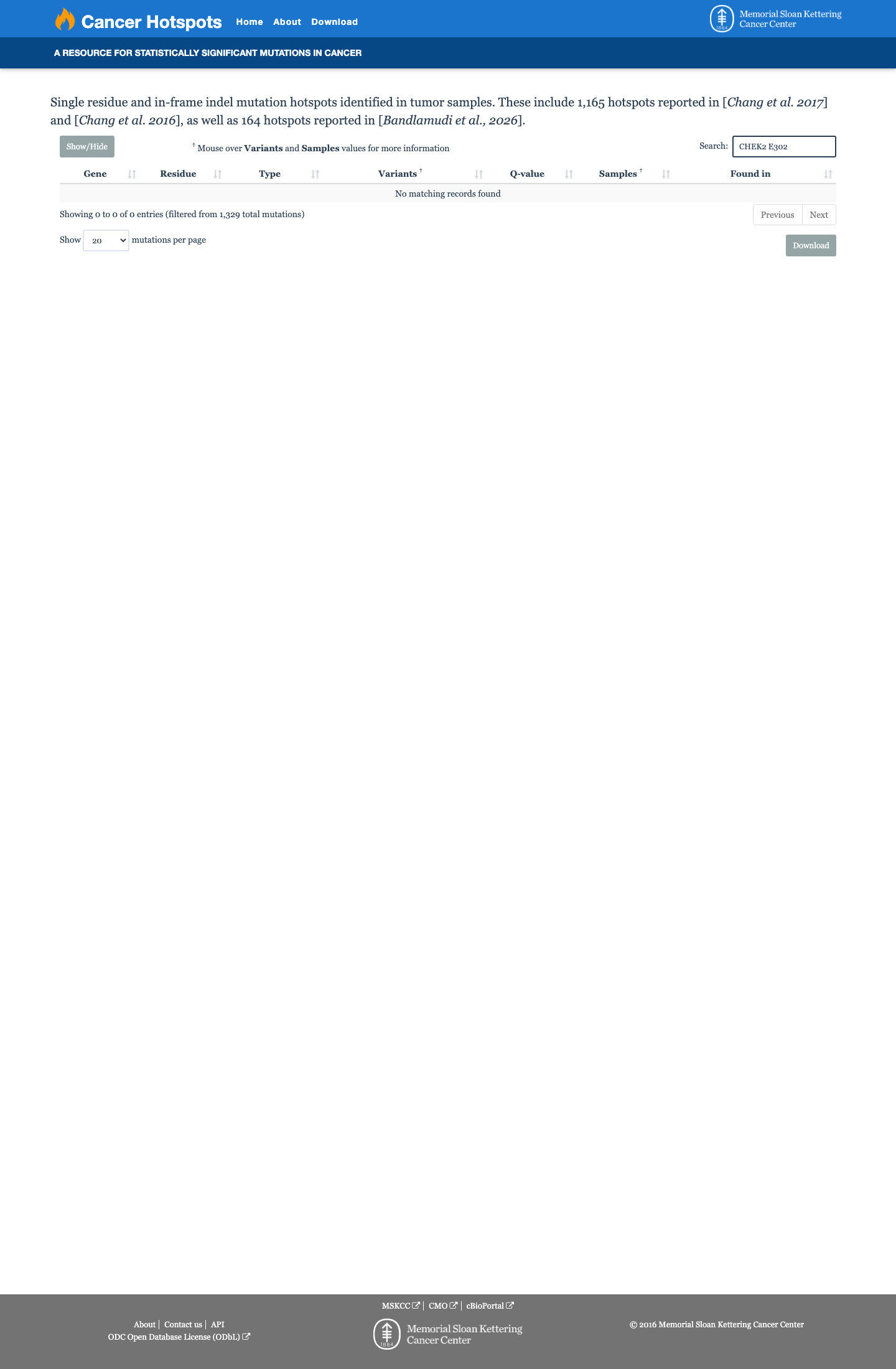
The CHEK2 c.906A>C (p.Glu302Asp, p.E302D) variant has not been observed in COSMIC and has been reported in ClinVar as a variant of uncertain significance.
clinvar ↗2
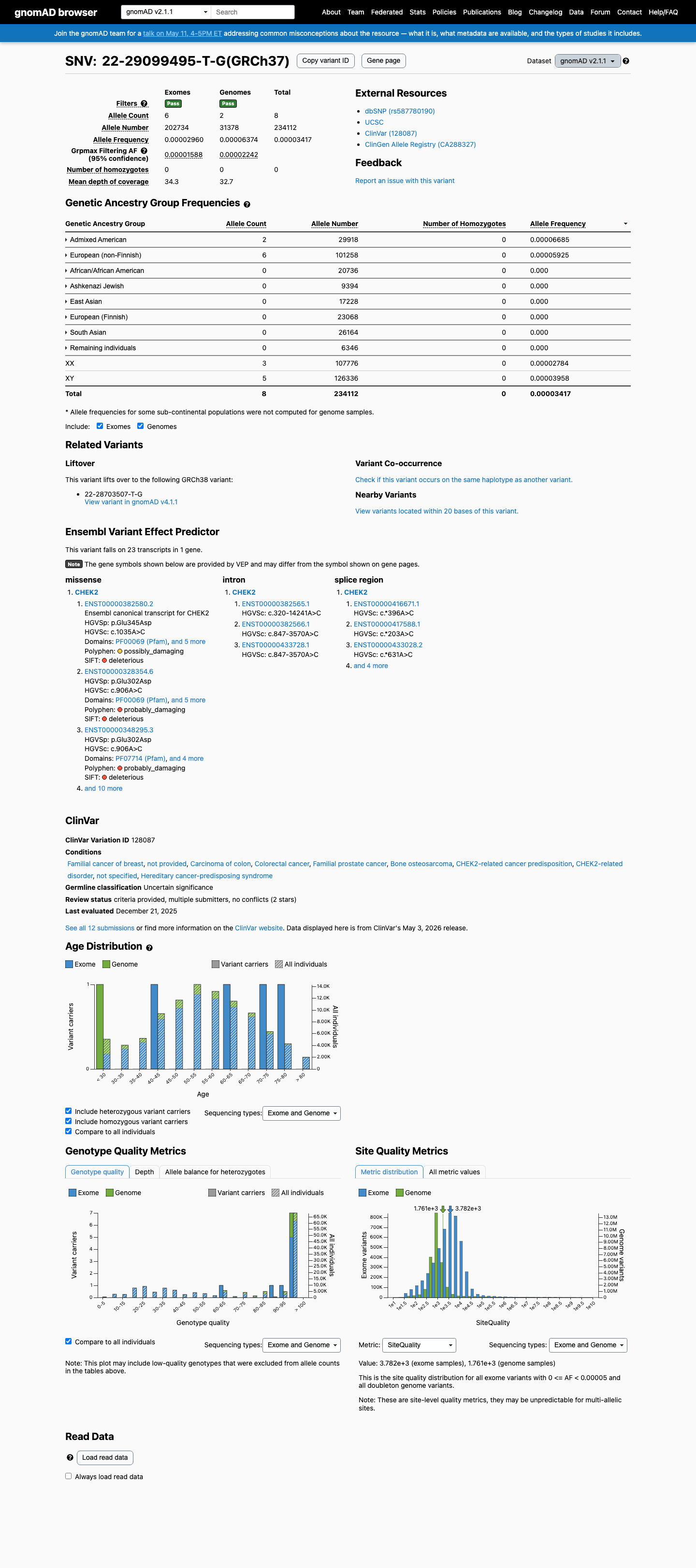
This variant is present at low frequency in population databases, with gnomAD v2.1 total allele frequency 0.00342% (8/234112) and gnomAD v4.1 total allele frequency 0.00097% (14/1437854); the highest observed subpopulation frequency is 0.03471% (2/5762) in Middle Eastern individuals, which remains below a 0.1% rarity threshold.
gnomad_v2 ↗ gnomad_v4 ↗3
In silico data do not show a consistent damaging signal: SpliceAI predicts no significant splice impact with a maximum delta score of 0.01, REVEL is 0.41, and BayesDel is 0.0235302.
spliceai ↗