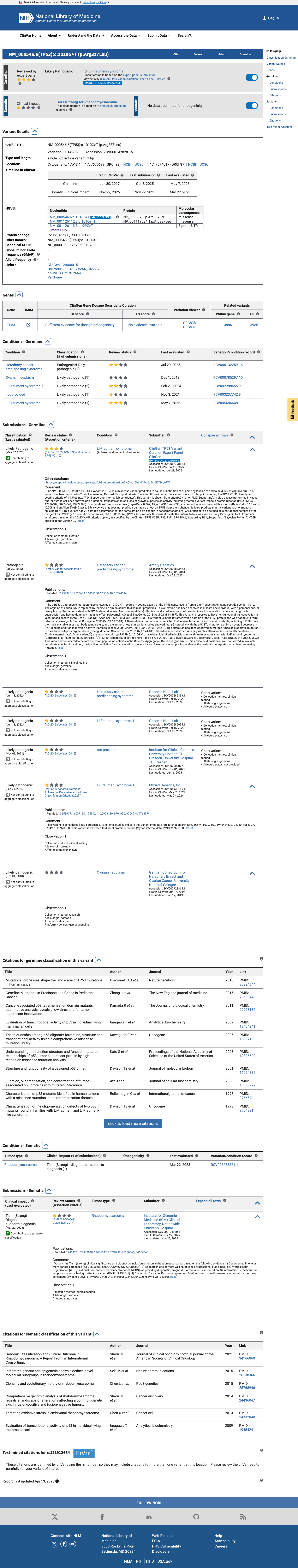
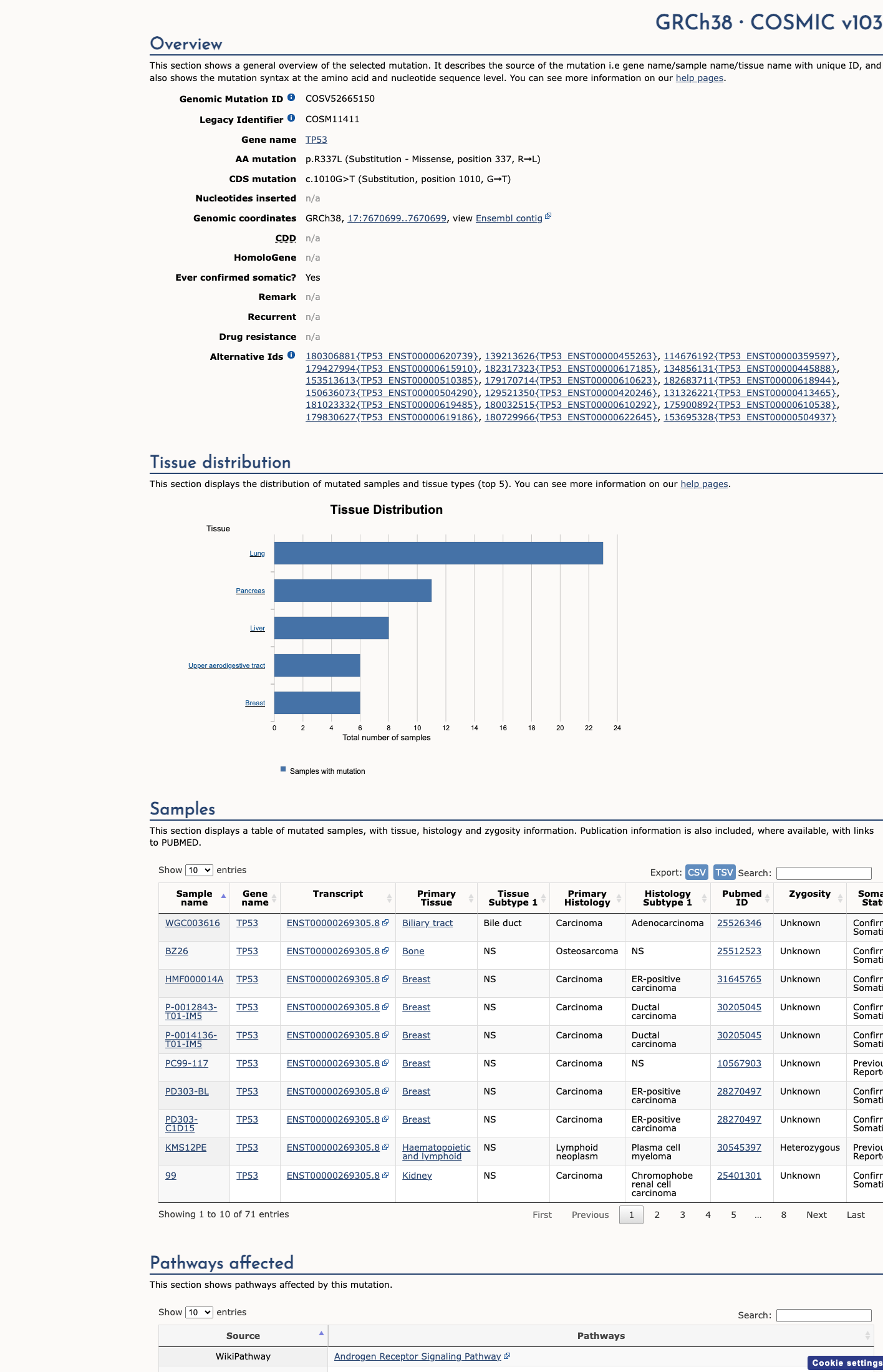
The TP53 c.1010G>T (p.Arg337Leu; p.R337L) variant has been observed in somatic cancers in COSMIC (COSV52665150, n=69) and has been reported in ClinVar, including a ClinGen TP53 expert panel classification of Likely Pathogenic.
clinvar ↗This variant is absent from gnomAD v2.1 and gnomAD v4.1, supporting rarity and meeting the TP53 PM2_Supporting threshold of less than 0.00003.
gnomad_v2 ↗ gnomad_v4 ↗ cspec ↗In TP53 functional data summarized by the TP53 VCEP, p.Arg337Leu was non-functional with loss-of-function findings and monomerization, supporting a damaging effect on protein function and meeting PS3.
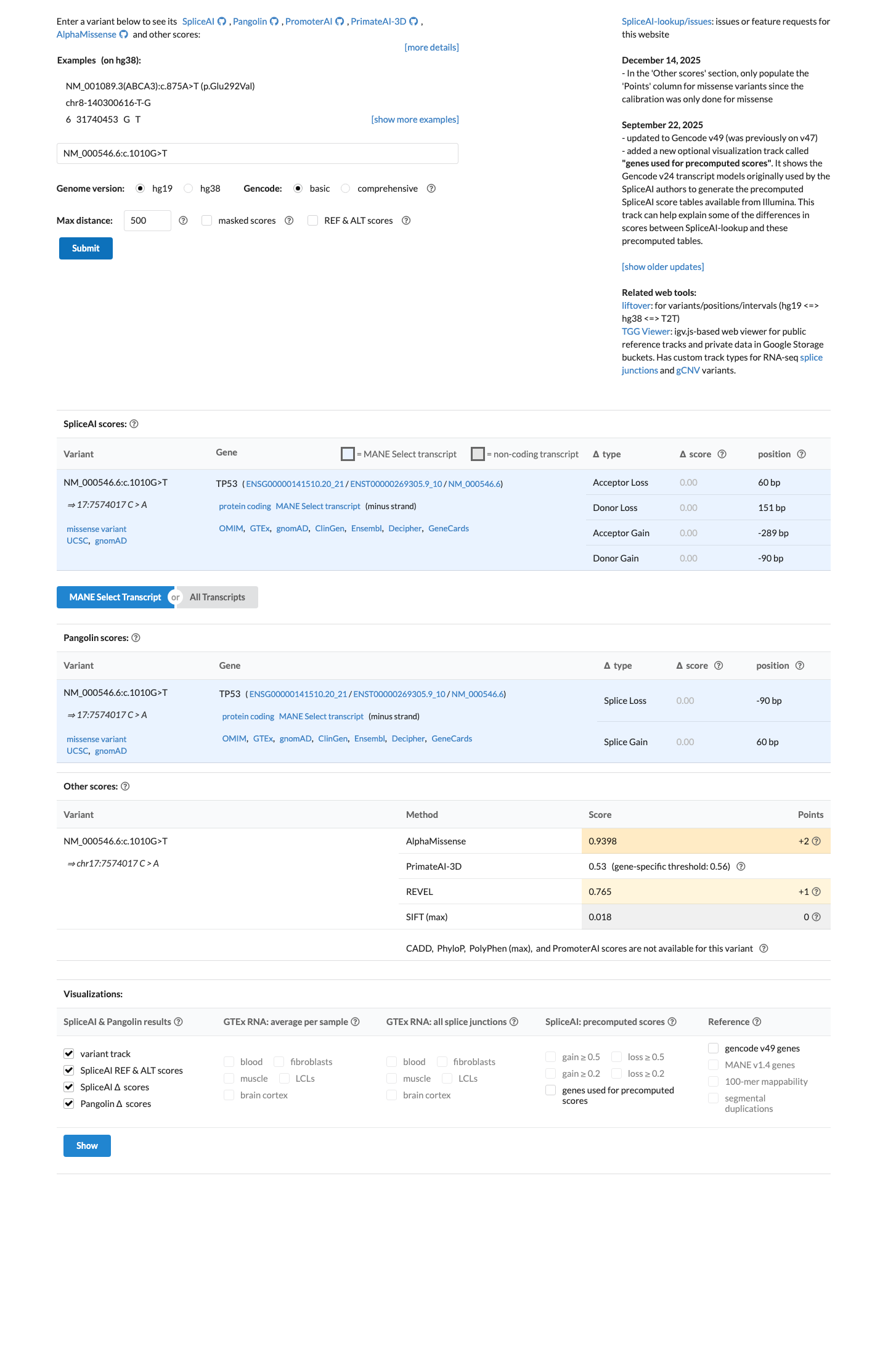
PMID:12826609 ↗ PMID:16007150 ↗ PMID:30224644 ↗In the TP53-specific computational framework, SpliceAI predicts no splice effect (max delta score 0.00) and the TP53 VCEP bioinformatic worksheet assigns BP4 with a BayesDel score of 0.0670081; REVEL was 0.765, but the TP53 precomputed PP3/BP4 assignment supports BP4 rather than PP3 for this variant.
spliceai ↗ cspec ↗