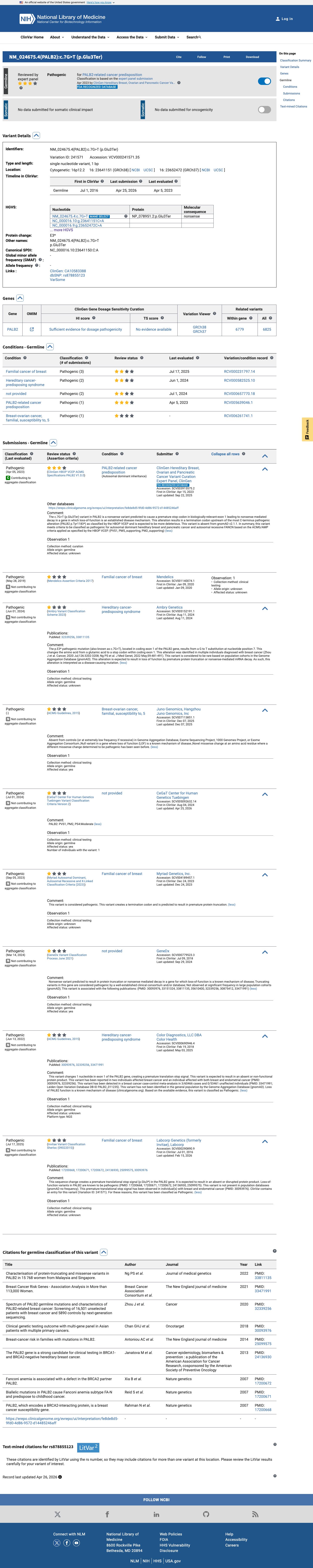
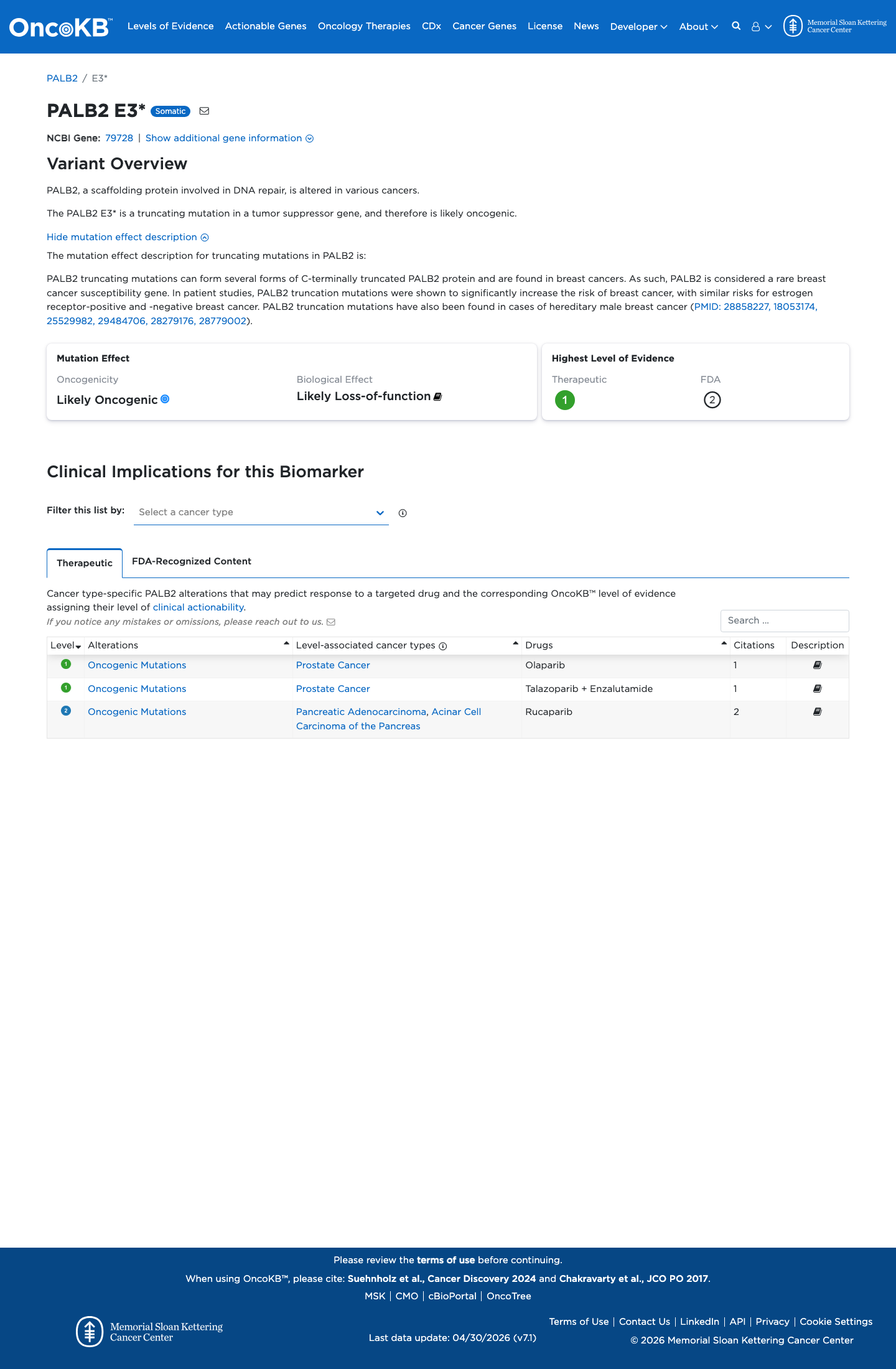
The PALB2 c.7G>T (p.Glu3Ter, p.E3*) variant has not been observed in somatic cancers in COSMIC and has been reported in ClinVar as pathogenic, including review by the ClinGen Hereditary Breast, Ovarian and Pancreatic Cancer expert panel.
clinvar ↗This variant is absent from gnomAD v2.1 and gnomAD v4.1, placing it below the PALB2 PM2_Supporting population threshold of 0.000333%.
gnomad_v2 ↗ gnomad_v4 ↗ cspec ↗This variant introduces a premature stop codon at amino acid 3, and the active PALB2 expert specification supports loss of function as an established disease mechanism, consistent with application of PVS1 and the PALB2 truncation-based PM5 rule.
cspec ↗SpliceAI predicts a limited splice effect with a maximum delta score of 0.15, which is below the PALB2 PP3 threshold of 0.2 and above the BP4 threshold of 0.1, so computational splice evidence is indeterminate; a BayesDel score of 0.63 is available but is not the governing PP3/BP4 rule in this gene-specific framework.
spliceai ↗ cspec ↗