Classification rationale
1
The BRCA2 c.2259T>C (p.Phe753=) variant has been reported in ClinVar with only likely benign submissions.
clinvar ↗2
This variant is absent from gnomAD v2.1 and present in gnomAD v4.1 at 3/1614118 alleles (AF 1.8586e-06; grpmax FAF 6.8e-07), which is far below ENIGMA BRCA2 BA1 and BS1 thresholds but does not satisfy absence from controls for PM2.
gnomad_v2 ↗ gnomad_v4 ↗ cspec ↗3
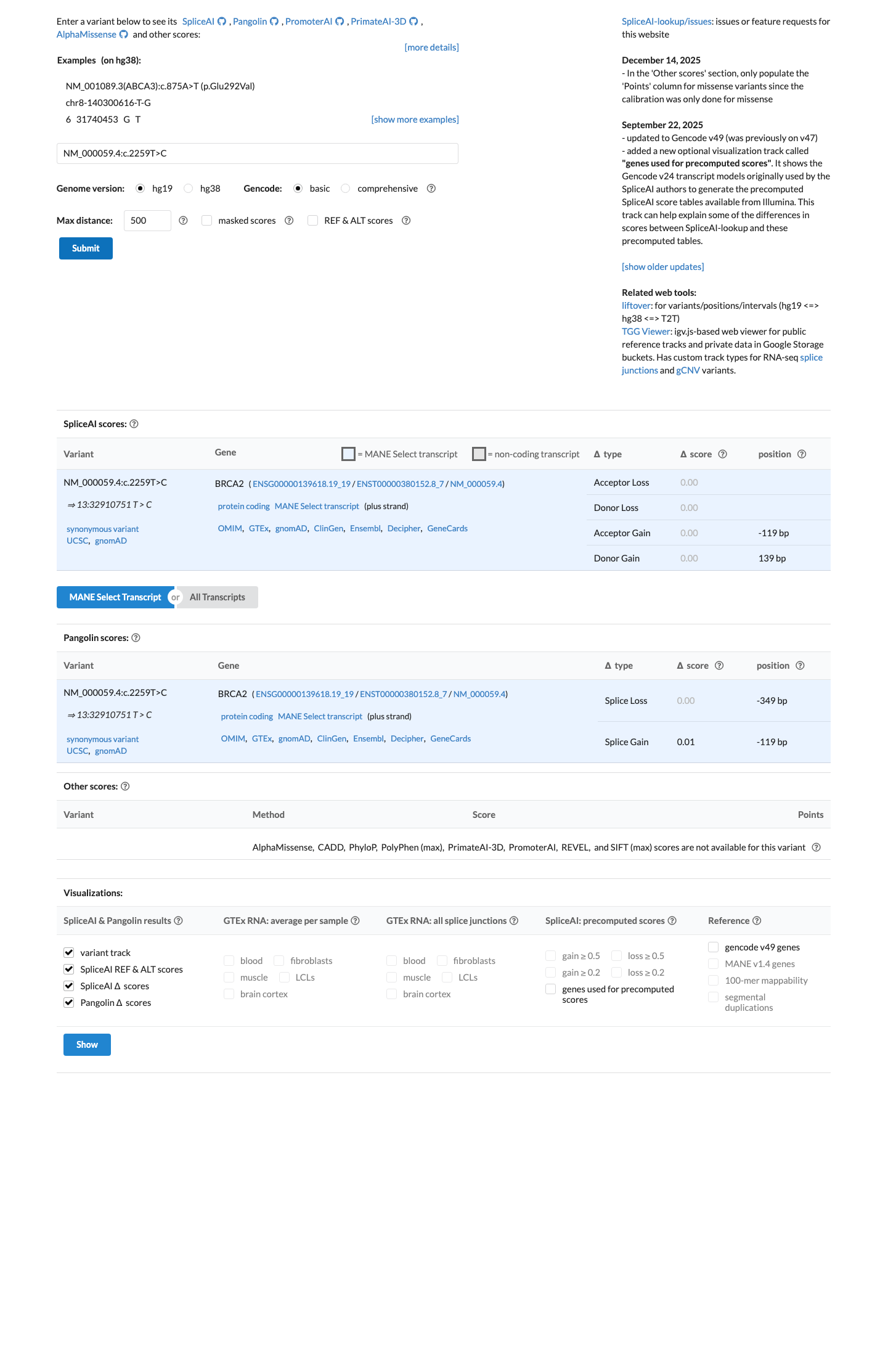
SpliceAI predicts no meaningful splice effect for this synonymous change (max delta score 0.00), and codon 753 lies outside the ENIGMA-defined BRCA2 clinically important domains, supporting BP1_Strong and not supporting PP3 or PVS1.
spliceai ↗ cspec ↗