Classification rationale
1
The NF1 c.369C>G (p.Thr123=) variant has been reported in ClinVar, where benign and likely benign classifications predominate.
clinvar ↗2
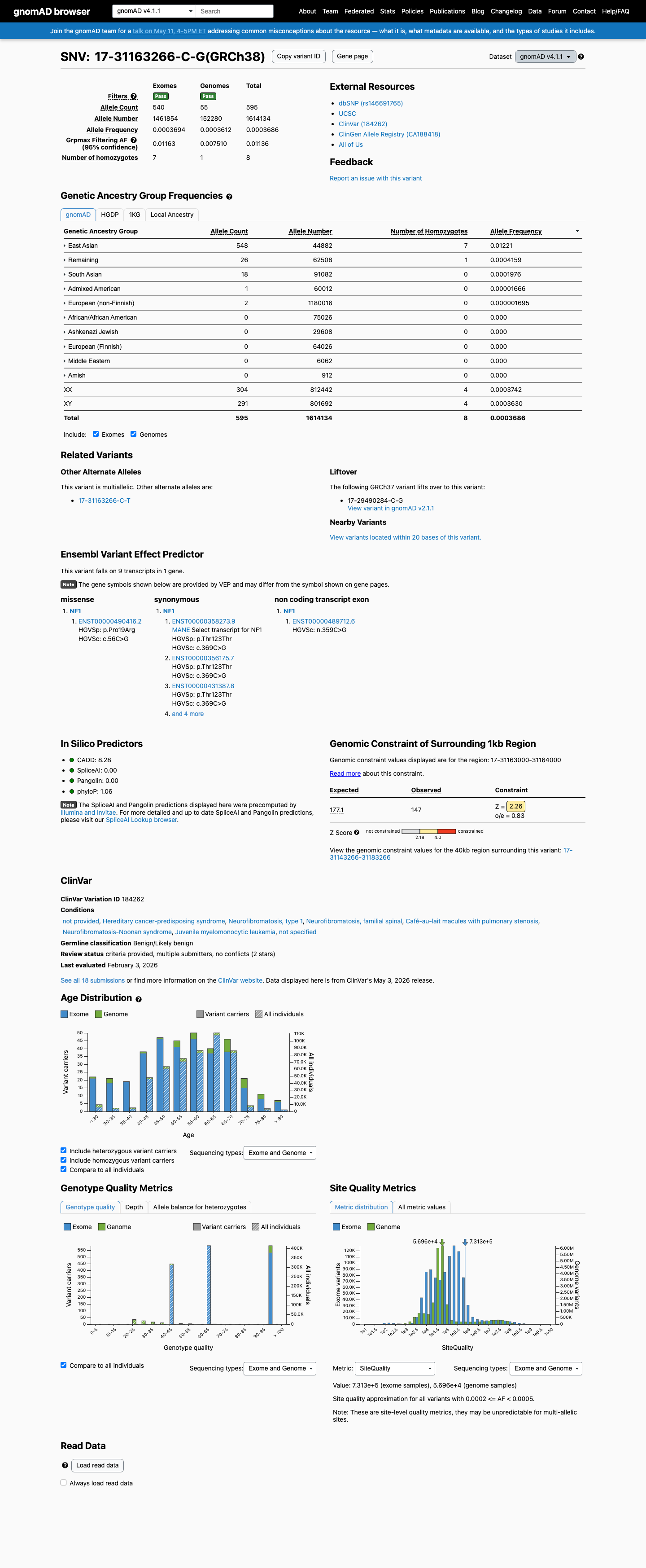
This variant is present in population databases at high frequency, including East Asian allele frequencies of 1.11791% in gnomAD v2.1 and 1.22098% in gnomAD v4.1, supporting BS1 and arguing against rarity-based pathogenic criteria.
gnomad_v2 ↗ gnomad_v4 ↗3
This synonymous variant is predicted to have no significant splice impact by SpliceAI, with a maximum delta score of 0.00, supporting BP7 and not supporting PP3.
spliceai ↗