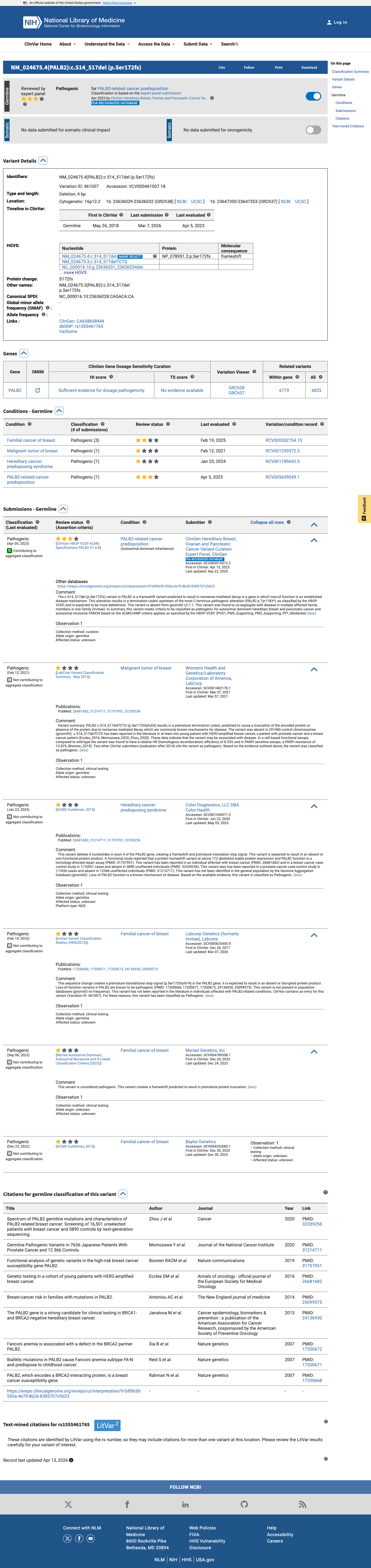
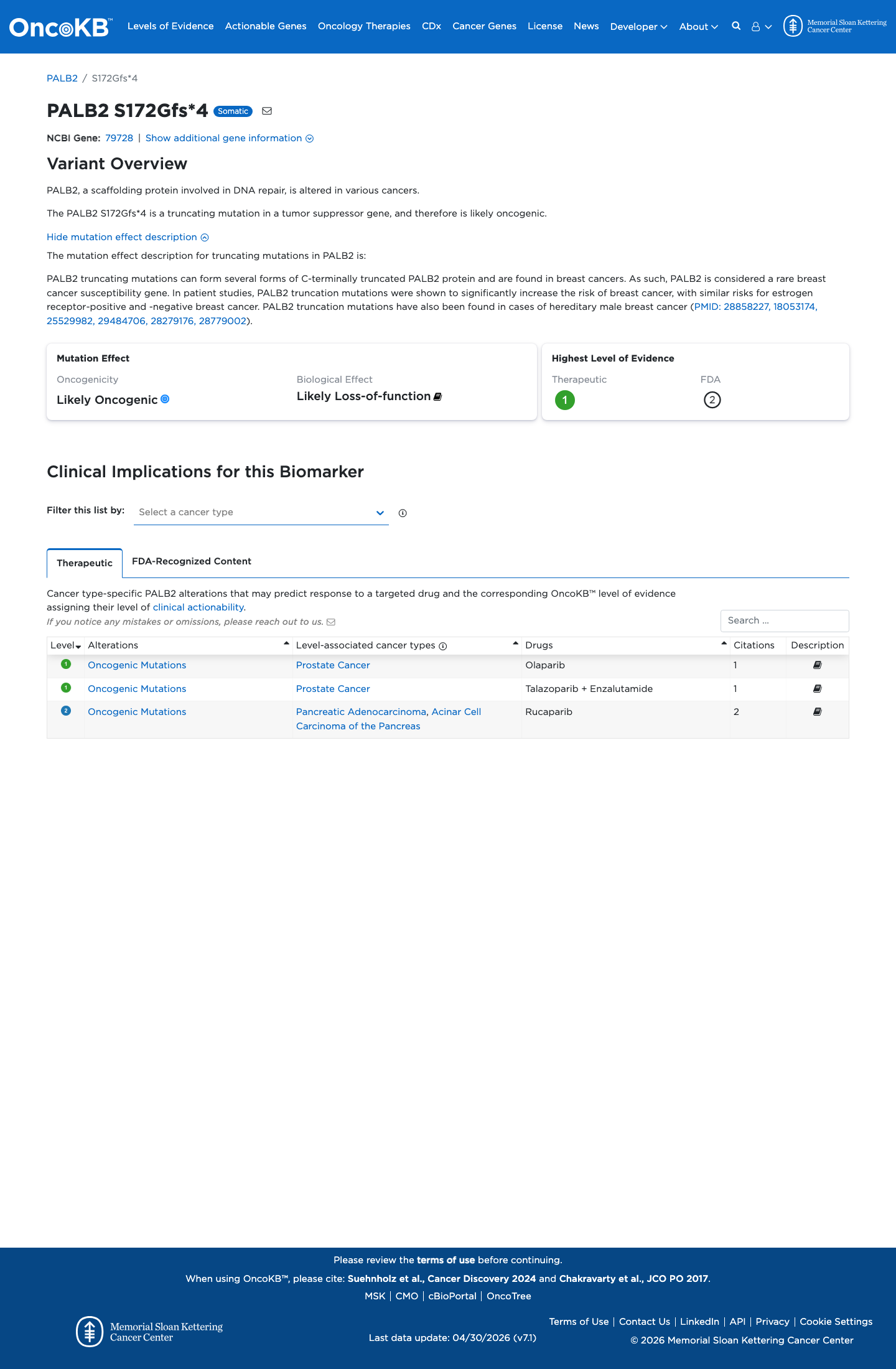
The PALB2 NM_024675.3:c.514_517del (NP_078951.2:p.(Ser172GlyfsTer4); p.(S172Gfs*4)) variant has not been observed in somatic cancers in COSMIC and has been reported in ClinVar as pathogenic, including expert panel review.
clinvar ↗This variant is absent from gnomAD v4.1 and gnomAD v2.1, placing its observed population frequency below the PALB2 PM2_Supporting threshold of 0.000333% and well below the BS1 and BA1 thresholds.
gnomad_v4 ↗ gnomad_v2 ↗ cspec ↗This deletion causes an early frameshift with a premature stop codon, and the PALB2 disease-specific framework supports loss-of-function interpretation for this null variant with PVS1_VeryStrong and PM5_Supporting for a truncating variant upstream of p.Tyr1183.
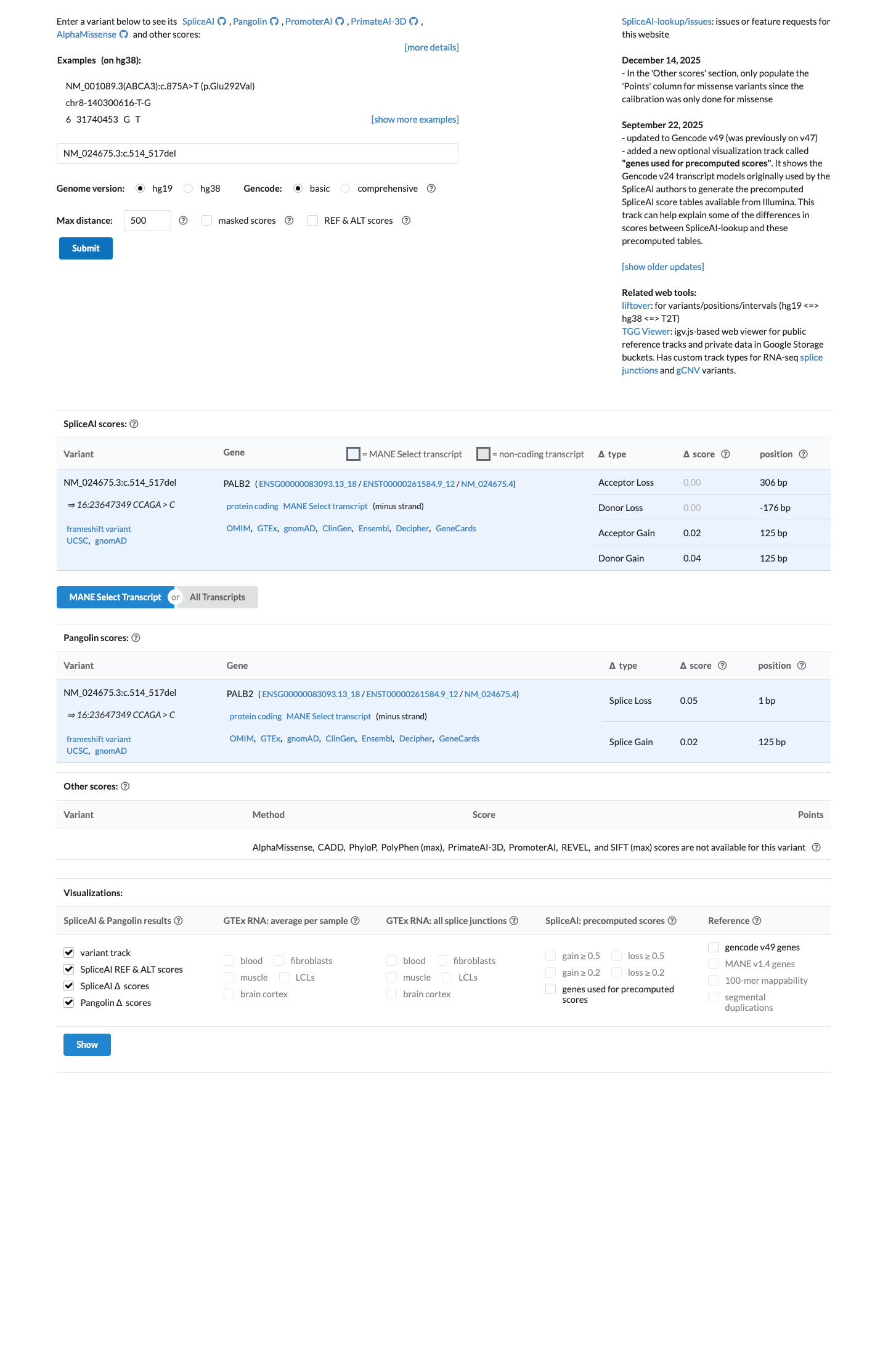
cspec ↗SpliceAI predicts no significant splice impact for this variant (max delta score 0.00), so PP3 is not met and the interpretation is driven by the truncating effect rather than an added predicted splice defect.
spliceai ↗ cspec ↗