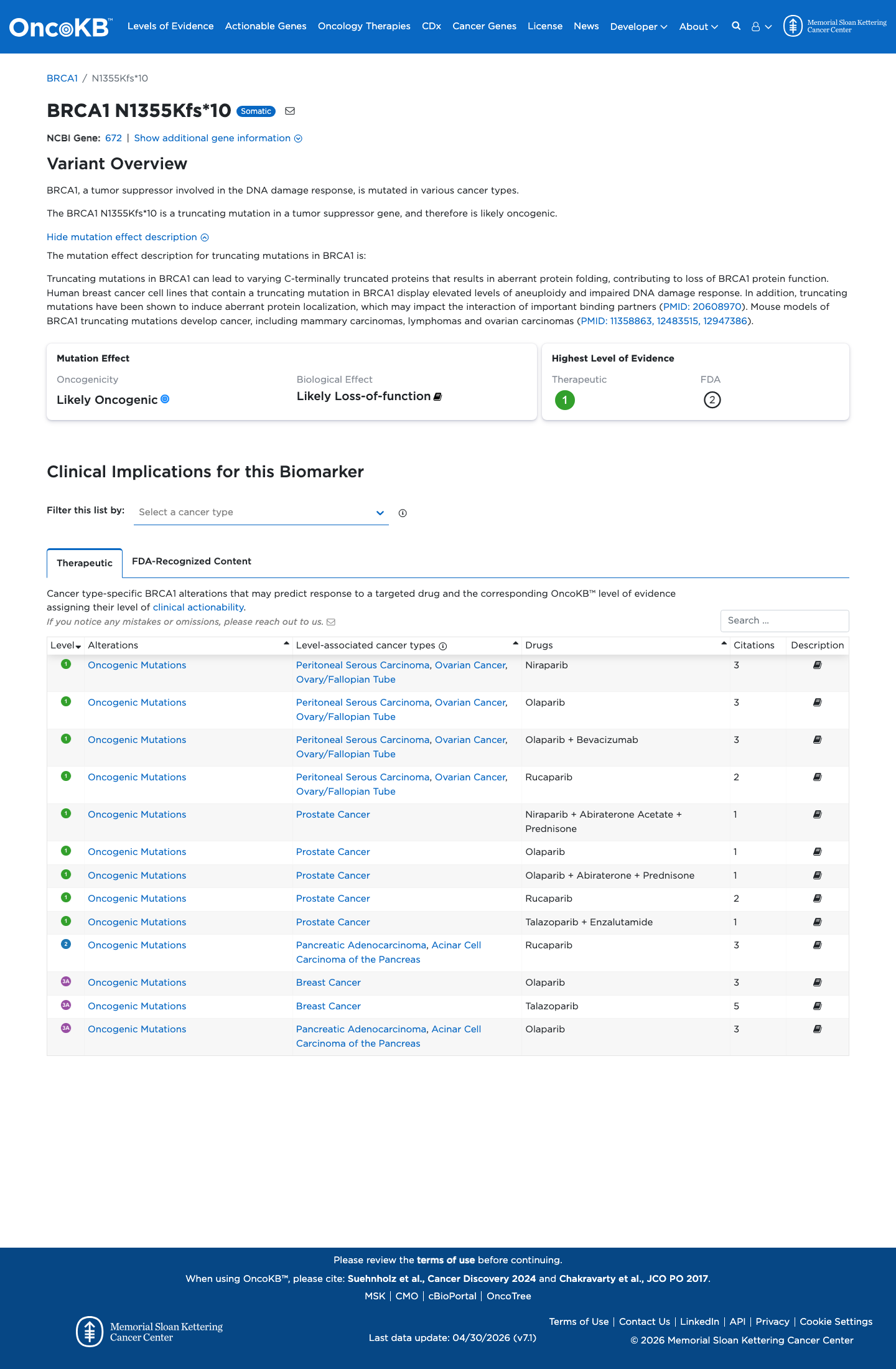
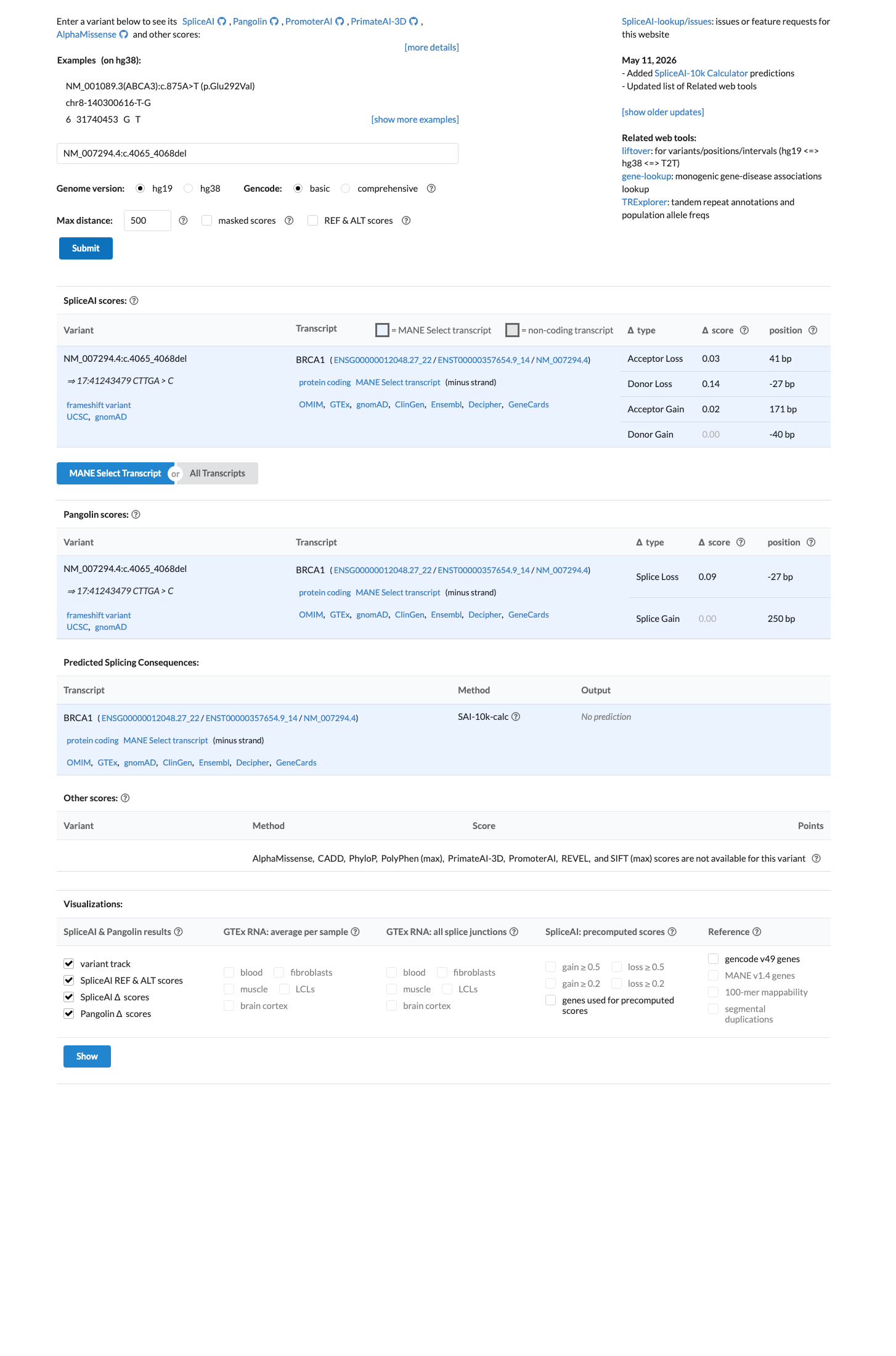
The BRCA1 NM_007294.4:c.4065_4068del (NP_009225.1:p.(Asn1355LysfsTer10), p.(N1355Kfs*10)) variant has been reported in ClinVar as Pathogenic by the ENIGMA expert panel and by multiple clinical laboratories.
clinvar ↗This variant is present at very low frequency in gnomAD v2.1 (3/250844 alleles; AF 1.19596e-05, 0.00120%; grpmax FAF 2.93e-06), which is below the BRCA1 BA1 and BS1 thresholds but means the variant is not absent from controls, so PM2 is not met.
gnomad_v2 ↗ cspec ↗The BRCA1 clinical-history likelihood-ratio dataset lists this deletion as c.4065_4068delTCAA in 37 probands with LR 6590.9969, supporting pathogenic clinical-history evidence under the ENIGMA BRCA1 framework.
PMID:31853058 ↗This frameshift lies in BRCA1 exon 10(11) and predicts a premature stop codon at residue 1364; the ENIGMA BRCA1 exon-specific Table 4 supports PVS1 and PM5_Strong (PTC) for truncating variants in this exon.
cspec ↗