Classification rationale
1
2
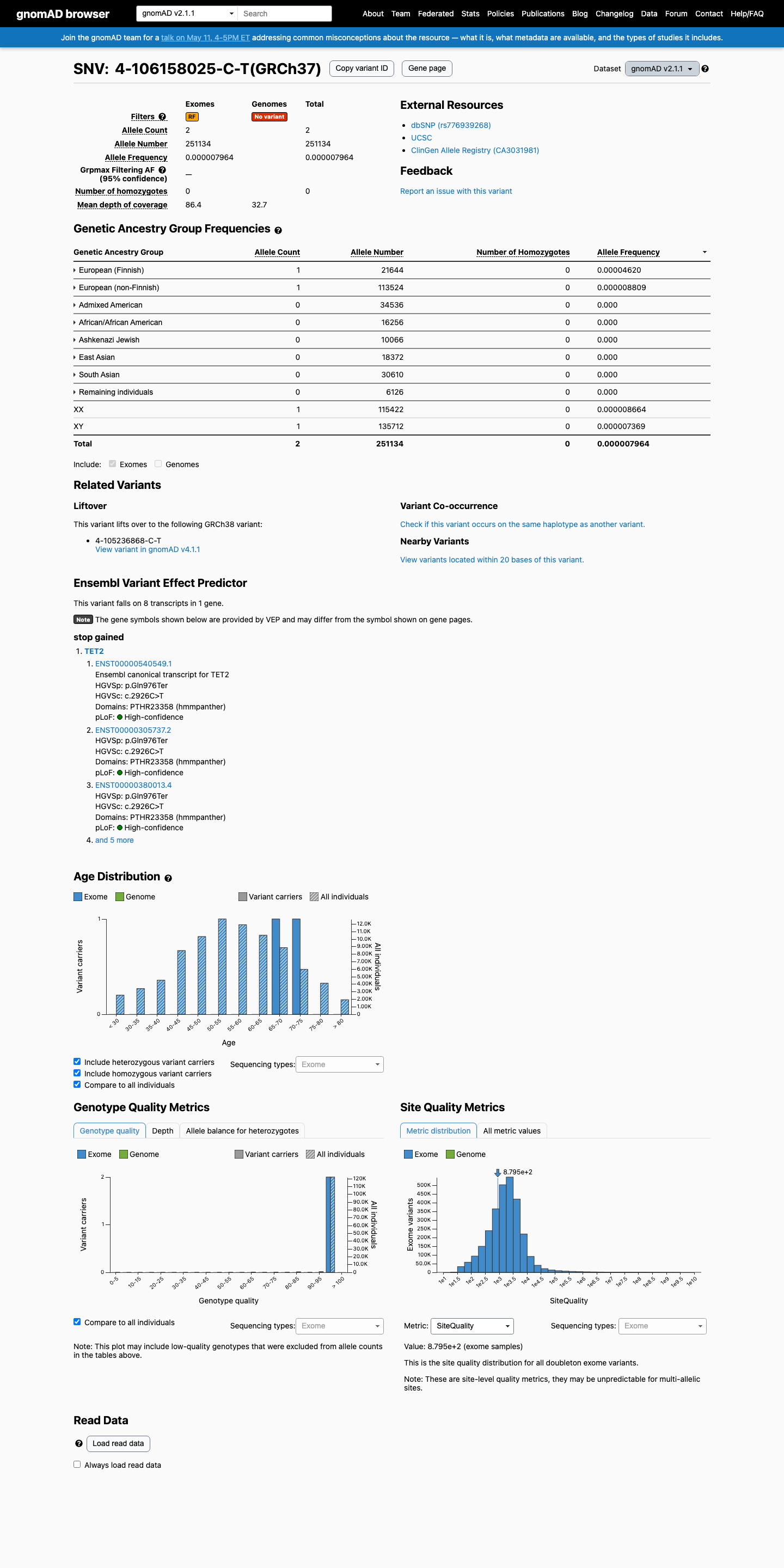
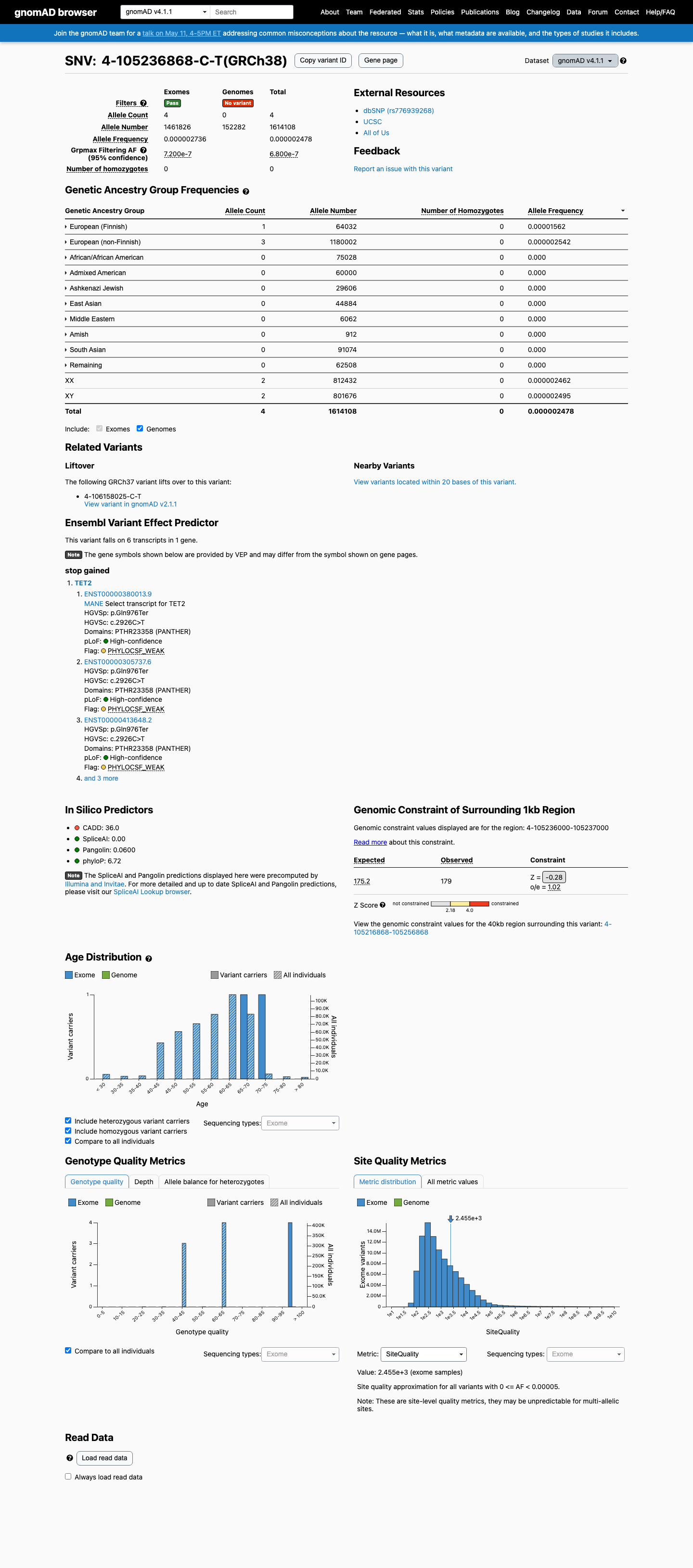
This variant is present at very low frequency in population databases, with allele frequency 7.96388e-06 in gnomAD v2.1 and 2.47815e-06 in gnomAD v4.1, both below the PM2 threshold.
gnomad_v2 ↗ gnomad_v4 ↗3
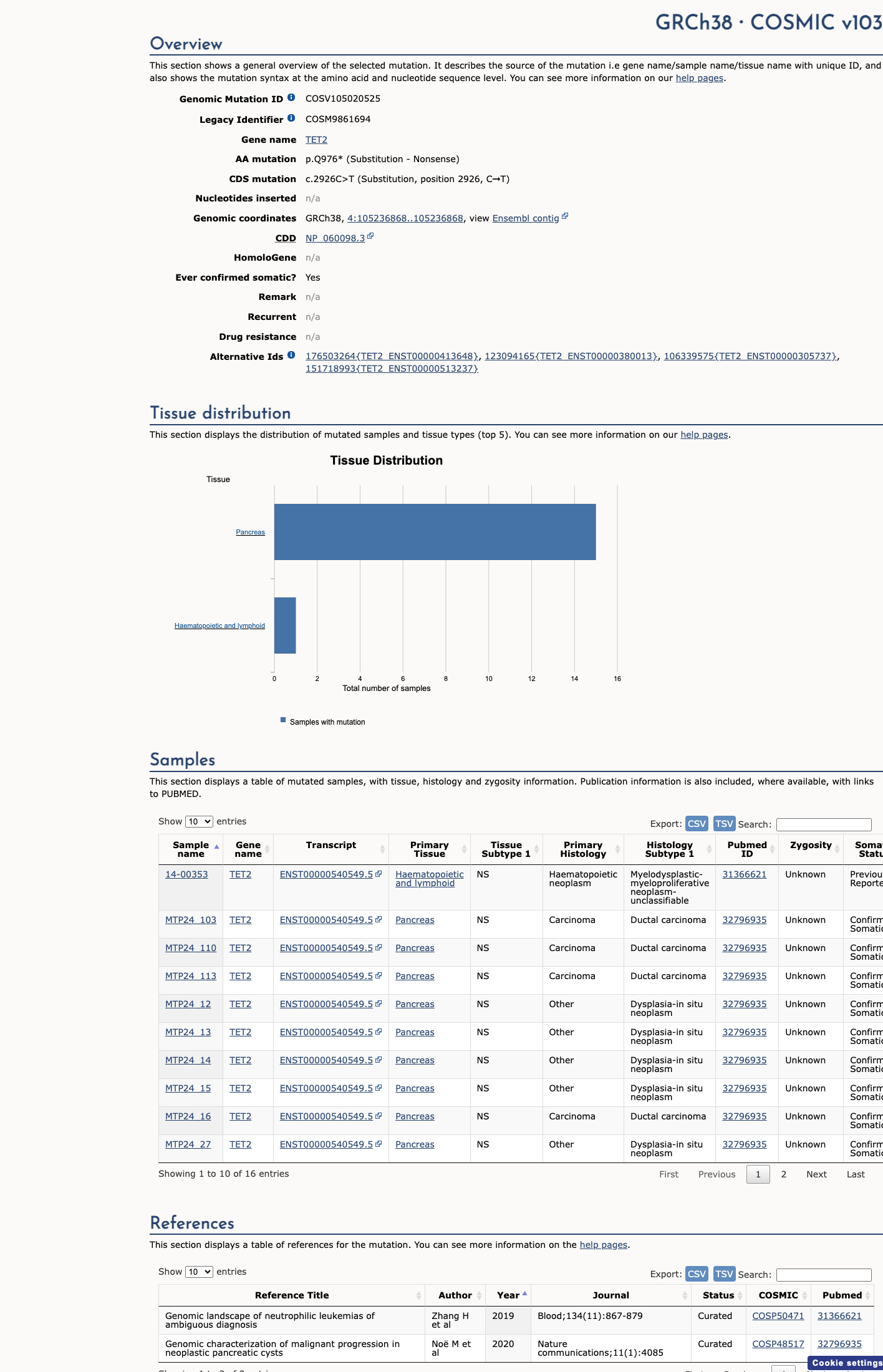
Available gene-level studies support TET2 loss-of-function biology and structural importance of the C-terminal catalytic region, but no variant-specific functional assay for p.(Gln976Ter) was identified.
PMID:21057493 ↗ PMID:24315485 ↗ oncokb ↗4
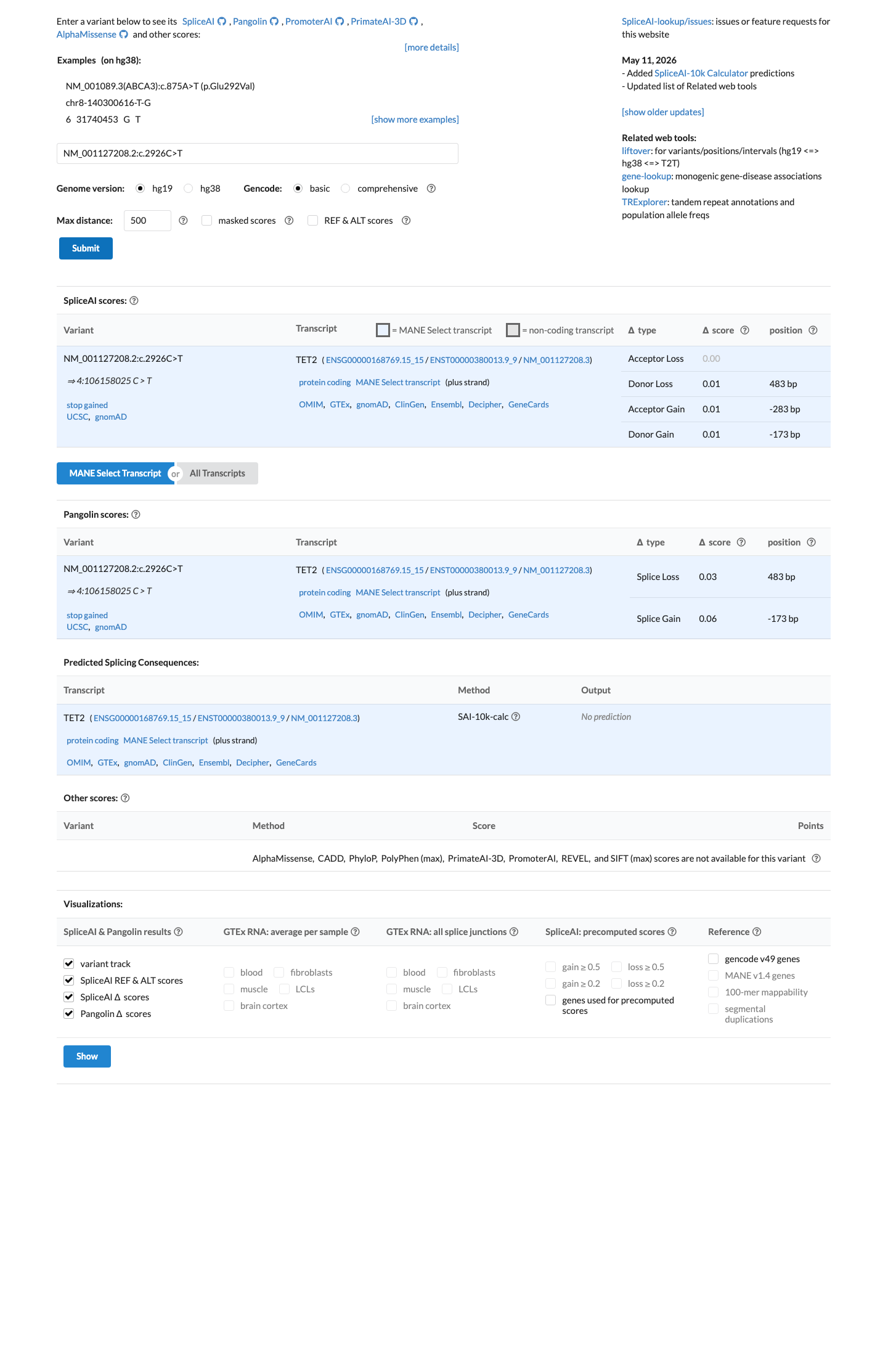
Computational review showed no significant predicted splice effect by SpliceAI, with a maximum delta score of 0.01; REVEL was unavailable, and BayesDel score 0.294732 did not provide sufficient support for PP3 or BP4 for this nonsense variant.
spliceai ↗