Classification rationale
1
The ZRSR2 c.617T>C (p.Phe206Ser; p.F206S) variant has not been reported in ClinVar and does not lie in a statistically significant hotspot.
clinvar ↗ hotspots ↗2
This variant is absent from gnomAD v2.1 and gnomAD v4.1, corresponding to an observed population frequency of 0%, which is below the 0.1% PM2 threshold.
gnomad_v2 ↗ gnomad_v4 ↗3
Available curated review did not identify variant-specific reviewed functional evidence for this variant.
oncokb ↗4
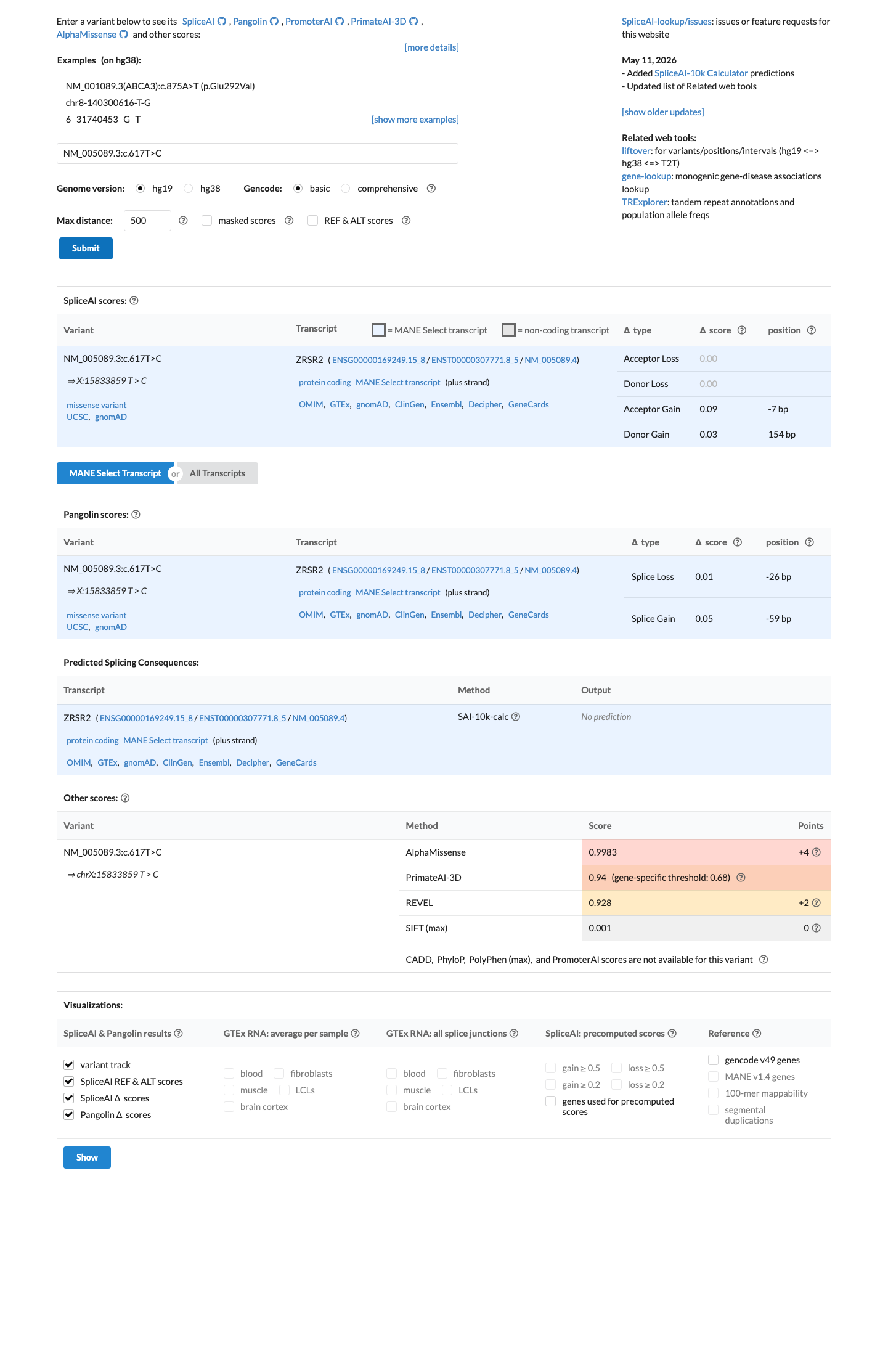
In silico evidence is mixed: BayesDel is positive at 0.563308, while SpliceAI predicts no significant splice impact with a maximum delta score of 0.09, so computational findings do not currently support PP3 or BP4.
spliceai ↗