Classification rationale
1
2
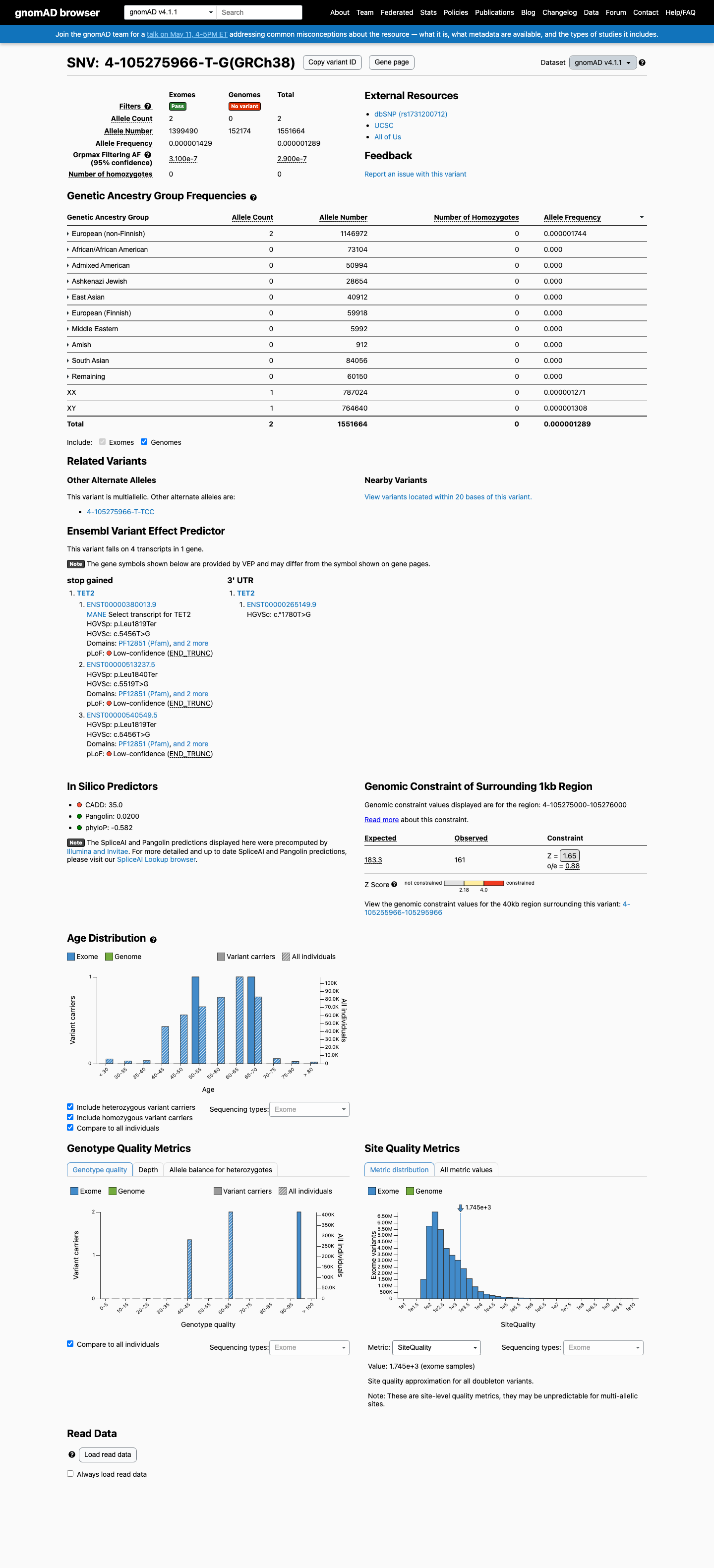
This variant is absent from gnomAD v2.1 and present in gnomAD v4.1 at 2/1,551,664 alleles (AF 0.00013%; highest population AF 0.00017%), which is below the 0.1% PM2 threshold and far below benign-frequency thresholds.
gnomad_v2 ↗ gnomad_v4 ↗3
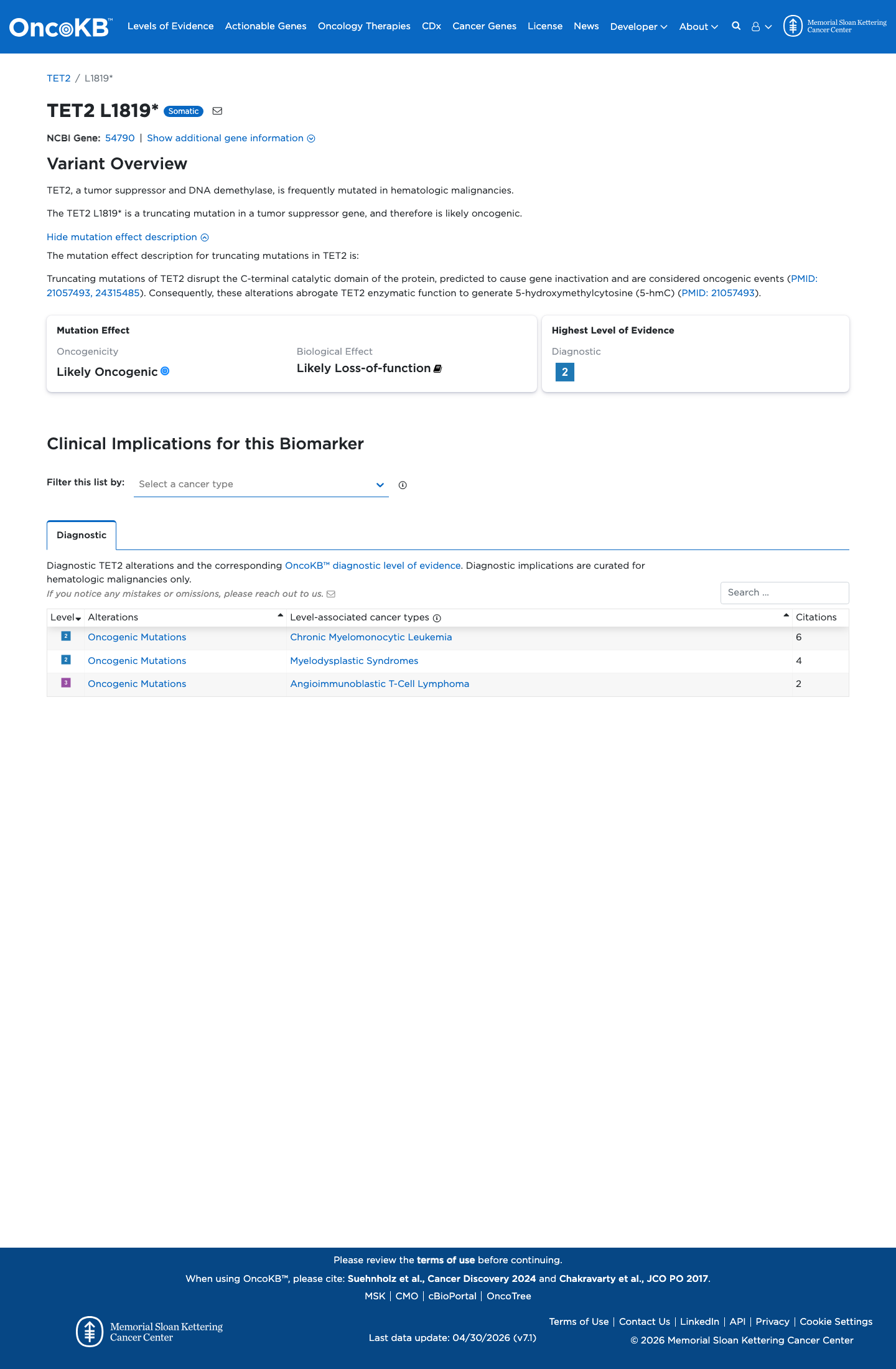
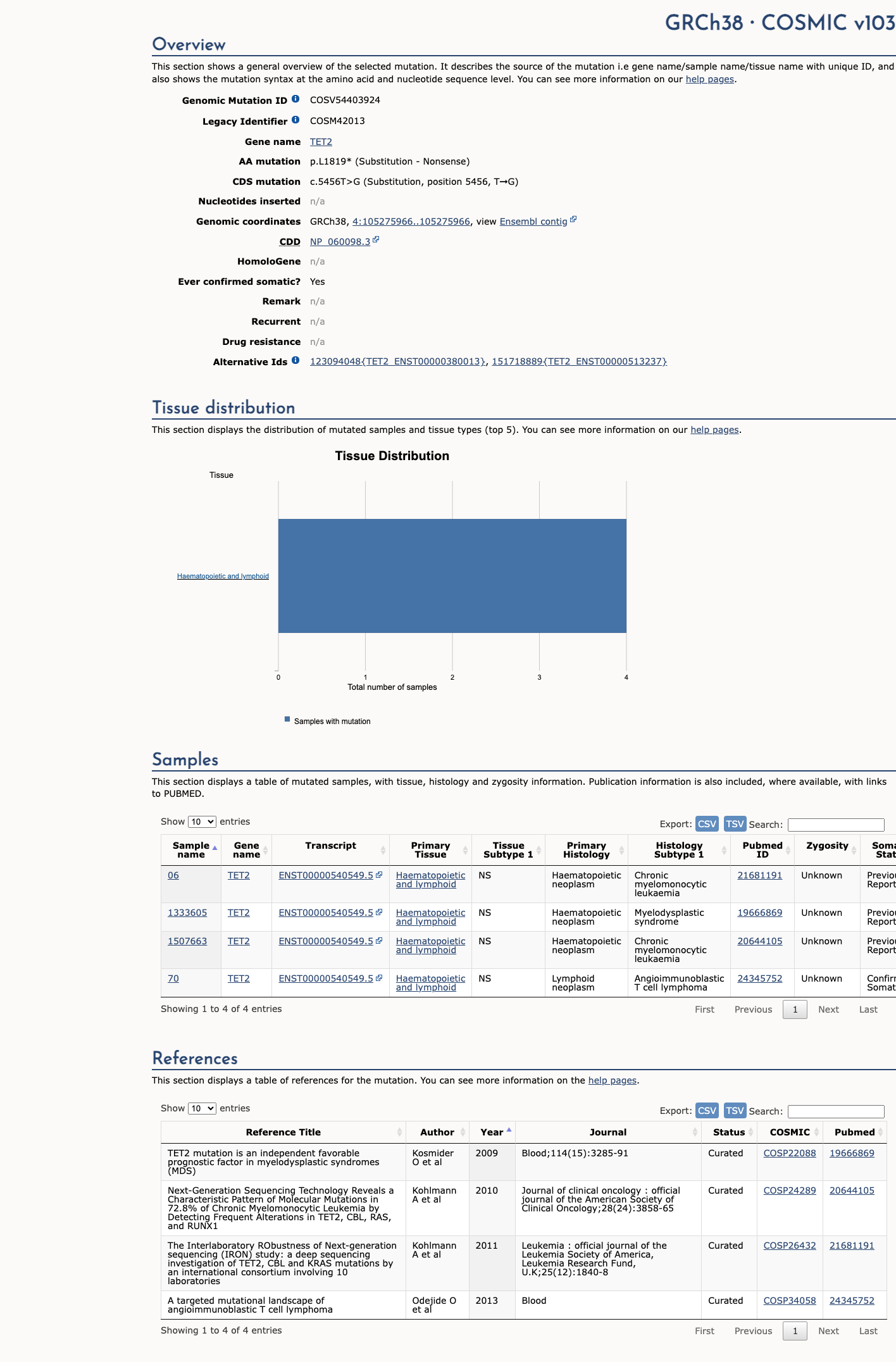
Published TET2 studies support the biological importance of TET2 loss of function, but no variant-specific functional assay for p.(Leu1819Ter) was identified.
oncokb ↗ PMID:21057493 ↗ PMID:24315485 ↗4
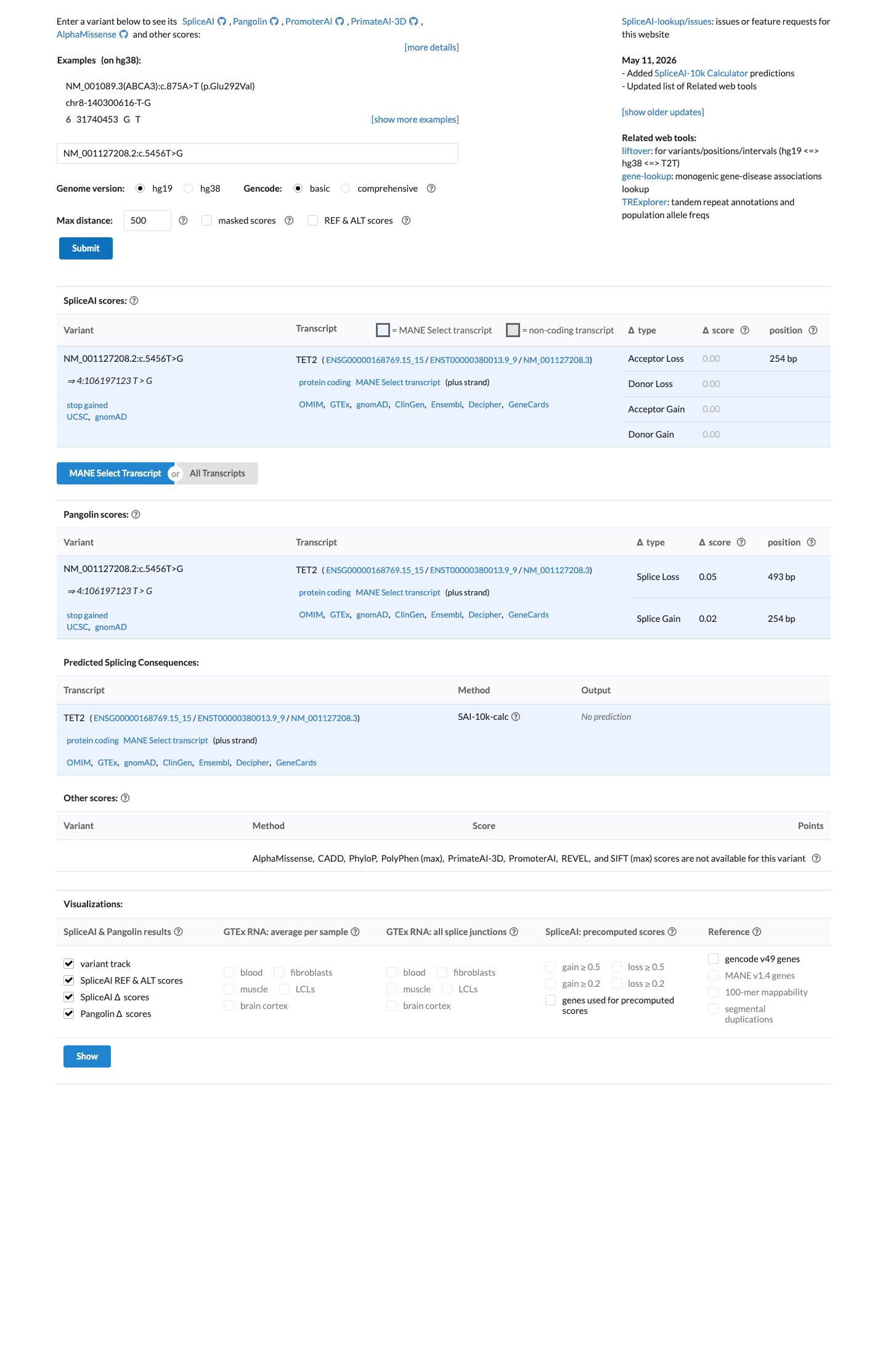
This nonsense change occurs in the last exon, SpliceAI predicts no significant splice impact (max delta score 0.00), REVEL was unavailable, and BayesDel score 0.288612 does not independently resolve pathogenicity for this variant.
spliceai ↗